| HE, SNGH esterase domain | |
|---|---|
| Identifiers | |
| Symbol | ? |
| InterPro | IPR007142 |
| HE, Hemagglutinin domain | |
|---|---|
| Identifiers | |
| Symbol | ? |
| InterPro | IPR003860 |
Hemagglutinin esterase (HEs) is a glycoprotein that certain enveloped viruses possess and use as an invading mechanism. HEs helps in the attachment and destruction of certain sialic acid receptors that are found on the host cell surface.[1] Viruses that possess HEs include influenza C virus, toroviruses, and coronaviruses of the subgenus Embecovirus (which does not include SARS-like coronaviruses). HEs is a dimer transmembrane protein consisting of two monomers, each monomer is made of three domains. The three domains are: membrane fusion, esterase, and receptor binding domains.
The different HEs enzyme activities include: receptor binding activity, receptor hydrolysis (esterase) activity, and membrane fusion activity. The receptor binding activity involve the attachment of HEs to N-acetyl-9-O-acetylneuraminic acid (9-O-Ac- Neu5Ac) of glycolipids and glycoproteins and in turn serve as viral receptor.[2] Receptor hydrolysis (esterase) activity allows virus particles to escape the infected cell by removing an acetyl group from the C9 position of terminal 9-O-Ac-Neu5Ac residues.[2] Membrane fusion activity helps in incorporation viral genome into the host cell cytoplasm by enhancing the attachment between the viral envelope and host cell membrane.
In certain influenza viruses, the cell surface consists of both hemagglutinin (HA) and neuraminidase (NA) proteins that encompass enzymatic activities, whereas hemagglutinin-esterase fusion (HEF) proteins have been found to be the primary single spike protein that combines all of the enzymatic activities listed above. HEF proteins have been tested to be high-temperature and low-pH resistant and are the primary source of virulence in viruses.[3] Influenza C have been shown to have unique HEF structure proteins that enhance its ability to infect the host cell compared to influenza A and B.
The folding of different domains in the hemagglutinin-esterase protein is important for intracellular transport of proteins from the endoplasmic reticulum to the Golgi apparatus. The presence of oligosaccharide chains in the E, F, and R domains of the HE enzyme also influence intracellular transport. Acylation of the hemagglutinin-esterase has shown to play an essential role in virus particle assembly replication. The exact process of enzyme catalytic cleavage has not yet been detailed out. However, proteolytic cleavage must occur before hemagglutinin-esterase membrane fusion activity. HEF proteins have a unique spikes hexagonal arrangement. This feature is unique to influenza C virus particles. The arrangement is a covering outside of the particle.
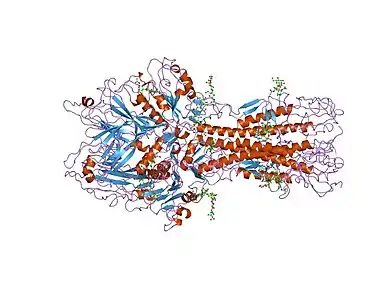
Structure
Certain studies revealed that coronavirus and toroviruses HE was originated from HEF glycoprotein that is found in influenza C viruses which resulted from alteration of hemagglutinin esterase from a trimer into a dimer glycoprotein.[1] During this process, the receptor destroying enzyme acetyl esterase domain stayed unchanged. However, the HE receptor binding domain has been altered in which that the ligand is bound in opposite orientation than before.[1] Both coronavirus and toroviruses HE monomers are made up of the same three domains: central esterase/hydrolase domain, receptor binding lectin domain, and membrane proximal domain which is small.[4] The two monomers of HE dimer in both CoV and ToV involve the same two contact regions (CR 1 and 2). CR 1 contain the receptor binding domain and contact region 2 that contain membrane proximal domain. Yet, ToV HE contacts region 2 contain additional esterase domain. As a result, the CR 2 surface is larger in ToV HEs than in CoV HEs. However, close to the carboxylic terminal membrane anchor, there are number of disulfide bridges between Cys385 of coronavirus HE that in turn keep the HE dimers connected to each other.[4]
In CoV HE, the two R domain beta sheets are connected to each other forming a continuous intermolecular beta sheet across the dimer interface. On the other hand, in ToV they are oriented at angles. As a result, the beta sheet of receptor binding domain in ToV is more twisted, the contact region 1 is smaller, and the R domains position are shifted along the Beta strands compared to CoV.[4]
Crystalline structure
"Initial studies using electron microscopy showed that the HEF spike forms a mushroom-shaped trimer consisting of a membrane-near stalk and a globular head".[2]
Later studies were able to examine and show a higher resolution structure (4.5 Å) of the hemagglutinin esterase fusion trimer using X-ray crystallography of the bromelain-cleaved ectodomain. Both hemagglutinin and hemagglutinin esterase fusion protein are similar in terms of structure and the folding of individual segments. yet, only 12% amino acid are identical between HA and HEF. One significant difference between HE and HEF is the presence of an additional bulge in HEF globular domain (bottom part of the domain) which contains the esterase region. The receptor-binding region in both HA and HEF is found in the upper part of the domain and contain only HEF1 residues. The stalk is made of three 60 Å long α- helices that contain: all sequences of HEF2 sequence, and certain HEF1 residues which are N-terminal residues (1–40), and C-terminal residues (367–432).[2]
The crystalline structure shows that the way that HEF binds to 9-O-Ac- Neu5Ac is the same as the way HA binds to Neu5Ac. The binding parts include an α-helix, a loop and an extended strand. There are hydrogen bonds between the amino acids (Tyr127, Thr170, Gly172, Tyr227 and Arg292) and the hydroxyl-groups of the ligand, and other residues form the structural support of the receptor binding site. A unique hydrophobic pocket is present in the HEF binding site that in turn accommodates the acetyl methyl group.[2]
Activity
Receptor binding activity
Glycolipids and glycoproteins contain N-acetyl-9-O-acetylneuraminic acid (9-O-Ac- Neu5Ac) that serve as viral receptor in which HEF binds to. HEF can bind to its receptor whether or not 9-O-Ac-Neu5Ac is attached by an α-2,3 or α-2,6 linkage to the next galactosyl residue. However, host specificity can be affected by terminal N-acetylneuraminic acid (Neu5Ac) and the glycosidic linkage of Neu5Ac. Influenza C virus can recognize 9-O-Ac-Neu5Ac on the surface of different cells due to its unique receptor specificity.[2]
Receptor hydrolysis (esterase) activity
The receptor hydrolase activity of HEF aids in the release of virus particles from an infected cell using esterase enzyme that cleaves acetyl from the C9 position of terminal 9-O-Ac-Neu5Ac. The esterase activity of HEF which is part of serine hydrolase class includes a nucleophilic attack of the hydroxyl group (OH) of a serine amino acid, with the help of two other amino acids (histidine and aspartic acid), on the carbonyl group of the substrate. Basic histidine enhances the reactivity of serine by polarizing and deprotonating its hydroxyl group. Along with that, aspartic acid polarizes histidine.[2]
X-ray crystallography of the crystalline structure of HEF showed that serine 57, aspartic acid 352 and histidine 355 are the important amino acids for the esterase activity. Also, early studies showed that mutation in Ser57 and His355 residues can completely stop the esterase activity of HEF.[2]
Membrane fusion activity
The membrane fusion activity between, the viral envelope and endocytic vesicles of host cell, is important to help the virus inject their genome into the cytoplasm of the cell. In order to activate membrane fusion, Cleaving the precursor proteins HEF0 and HA0 into the subunits into the subunits HEF1 and HEF2, then exposing these proteins to acidic pH must be done prior.[2]
Acidic pH causes protonation of specific amino acids that initiate certain rearrangement of the proteins. The protonated amino acid is found to be histidine while its pKa matches the pH of endosome. Studies showed that there is about 0.7 difference in the pH value that trigger the membrane fusion activity from strain to strain of both influenza A and C.[2]
Conformational change in HEF structure that occur at low pH results in the separation of fusion peptide from its location at the lower part of the stalk and exposing the outer surface of the molecule, so it can be inserted into the endosomal membrane. Another conformational change occur which cause the bending of the ectodomain to push the fusion peptide toward the transmembrane region. As a result of that, the virus and endosomal membranes get closer, exchanging lipids with hemifusion. Then, opening of a fusion pore and eventually complete merger of both lipid bilayers.[2]
Folding and intracellular transport
The folding of the hemagglutinin esterase protein and the way that the domains of the protein assembles contribute the transport of membrane and secretory proteins from the endoplasmic reticulum to the Golgi apparatus. Researchers found that trimerization occurs at a point before exiting the ER.[5] The monomers of the HE protein are folded before assembling is possible. Before the hemagglutinin esterase can report to the Golgi, it must be extensively folded and assembled.
The structure of hemagglutinin-esterase contributes to the intracellular transport. The hemagglutinin-esterase (HE) glycoprotein of influenza C virus is composed of three domains: a stem domain active in membrane fusion (F), an acetylesterase domain (E), and a receptor-binding domain (R).[6] The protein contains eight N-linked glycosylation sites, four (positions 26, 395, 552, and 603) in the F domain, three (positions 61, 131, and 144) in the E domain, and one (position 189) in the R domain.[6] Oligosaccharide chains in the domains influence intracellular transport. A study showed that it was evident that glycosylation at the two sites in the F domain (positions 26 and 603), in addition to that in the E domain (position 144), is required for the HE molecule to be transported from the endoplasmic reticulum and that mutant HEs lacking one of these three sites failed to undergo the trimer assembly.[6] Oligosaccharides are needed to maintain esterase activity in the F and R domains. If any of the domains lack an oligosaccharide chain, cell surface expression will be affected. It was found that HE monomer have acetylesterase activity because they possessed full-enzyme activity despite lack of an oligosaccharide chain.[6] Oligosaccharide chains are important for intracellular transport, but not for fusion activity. Thus, oligosaccharide chains do not really promote membrane fusion.
S-acylation and RAFT-localization
Acylation of the hemagglutinin-esterase enzyme is necessary for virus replication of influenza C virus. It was found that recombinant virus lacking the acylation site of HEF could be rescued, but viral titers were reduced by one log relative to wild type Flu C.[2] The resulting virus particles have a regular protein composition and no changes in their morphology were obvious by electron microscopy, but their hemolytic activity is reduced indicating a defect in membrane fusion.[2] This is in comparison to several HA protein subtypes that showed similar results.
The hemagglutinin-esterase-fusion protein has co- and post-translational modification, such as N-glycosylation, disulfide bond formation, S-acylation and proteolytic cleavage into HEF1 and HEF2 subunits.[2] The HEF protein of influenza C virus has only one stearate attached to a transmembrane cysteine. Whereas HA of influenza A and B virus are associated with membrane rafts, cholesterol- and sphingolipid-enriched nanodomains of the plasma membrane, HEF is thought to localize to the bulk phase of the plasma membrane.[2]
Proteolytic cleavage
The binding and cleavage properties of the influenza C virions hemagglutinin-esterase (CHE) protein for 9-O-acetyl groups on sialic acids have been used in various assays using whole virions.[7]
Proteolytic cleavage must occur before any membrane fusion activity of HE because it enables the protein to become activated by low pH. HEF proteins from all influenza C virus strains contain a monobasic cleavage site and are in this respect similar to HAs from human, porcine, equine and low pathogenic avian influenza A viruses.[2] Polybasic cleavage sites that are present in HA of highly pathogenic avian influenza A viruses and processed by the ubiquitous protease furin are not found in any HEF protein. Consequently, replication of influenza C virus is limited to the site of virus infection, the respiratory tract.[2] Unlike other influenza viruses, influenza C virus does not spread to other tissues. Multiple replication cycles of influenza C virus in tissue culture are enabled by addition of trypsin, whereas embryonated eggs produce infectious virus with cleaved HEF.[2]
The enzyme catalyzing proteolytic cleavage of HEF has not been identified so far, but since both HA and HEF can be cleaved by trypsin at similar concentrations in vitro(5~20 µg/mL) it seems likely that they are also activated by the same enzymes inside cells.[2] It is very common that HA is compared to HEF in many contexts.
Regular arrangement of HE spikes in virus particles
The only spike of influenza C virus, the hemagglutinin‐esterase‐fusion glycoprotein (HEF) combines receptor binding, receptor hydrolysis and membrane fusion activities.[8] Like other hemagglutinating glycoproteins of influenza viruses HEF is S‐acylated, but only with stearic acid at a single cysteine located at the cytosol‐facing end of the transmembrane region.[8] This HE protein however, has spikes in its structural organization as well.
HEF trimers on the surfaces of both spherical and filamentous particles are arranged in a reticular structure that has been described to consist mainly of hexagons.[2] This feature is unique to influenza C virus particles. Even when HEF is removed from the membrane, the polymeric reticular structure that it originally had can still be seen. These results indicate that the hexagonal arrangement is an intrinsic feature of HEF and does not require other viral proteins such as M1 and that its formation likely involves lateral interaction between the ectodomains of HEF.[2] The formation of the spike arrangement in virus particles acts like a coat around the virus particle by creating and covering it. This is similar to the hydrophobic effect in lipid bilayer membranes where nonpolar molecules and in the interior.
Location of N-glycosylation sites
HEF N-glycosylation sites are located in figure 1. One sequon is located in HEF2 and six in HEF1. There are three in the globular head and 2 in the hinge region that connects the stalk with the head. The site at position 589 is not glycosylated because it is too close to the membrane-spanning region and cannot be accessed by the oligosaccharide transferase. Glycosylation is crucial for proper folding because it protects it from proteolytic degradation from the host cell and is important for the presentation of antigenic epitopes.[2]
In influenza C
The primary structure of HEF in influenza C contains 641 amino acids. It is a typical type 1 transmembrane protein with a short N-terminal, cleavable signal peptide, a long ectodomain, a transmembrane region and a very short cytoplasmic tail. HEF is composed of two subunits, HEF1 consisting of the N-terminal and HEF2 consisting of the transmembrane domain and the cytoplasmic tail. Electron microscopy analyzing the crystal structure of HEF showed that the spike of HEF forms a mushroom-shaped trimer consisting of a membrane-near stalk and a globular head. HEF contains only asparagine-linked carbohydrates which indicates that O-glycosylation does not occur. The location of the individual glycosylation sites in the crystal structure are located on seven of the eight highly conserved N-glycosylation sequons; one is located in the subunit, HEF2 and the other six are located on the subunit HEF1. Three sites are in the globular head and two are in the hinge region that connects the stalk with the head. There is a site at position 589 on the crystallized structure that is not glycosylated and this may be due close location to the membrane-spanning regions and cannot be accessed by oligosaccharide transferase. The positions of HA in influenza A are quite similar to influenza C by the majority of its carbohydrate positions being located in the larger subunit.[2]
Location of intramolecular disulfide bonds
In HEF1, 12/15 cysteine residues form 6 intrachain disulfide linkages that stabilizes the globular head domain. There are two cysteine residues, Cys373 and Cys399 that do not form disulfide linkages in the mature protein. They are located at the hinge that connects the globular head with the stalk region. The rest of the cysteine residues form interchain disulfide bonds with HEF in the ectodomain area, near the bottom of the trimer. These disulfide bonds in HEF2 allows the subunit to perform large conformational changes that catalyze membrane fusion.[2]
In influenza viruses
In influenza C, there are 15 cysteine residues in subunit HEF1, 12 of the residues form six intrachain disulfide links that stabilize the globular head domain. Two of the cysteine residues are not required for proper folding and function of HEF and/or they do not form a disulfide linkage in the mature protein located at the connection hinge. The remaining cysteine rescues forms an interchain disulfide bond with the only cysteine residue in the ectodomain of subunit HEF2. This residue is located at the bottom of the trimer. In comparison, influenza A, has similar disulfide bond distributions with one bond connecting HA1 with HA2, the majority are intrachain bonds. The rare occurrence of disulfide bonds in HEF2 and HA2 subunits allows these subunits to perform large conformational changes that catalyze membrane fusion.[2]
Co- and post-translation modification
During translocation of HEF into the lumen of the ER, the N-terminal signal peptide is cleaved, and carbohydrates are attached. Disulfide bond linkages are formed and remodeled. These modifications affect the folding and trimerization of the molecule. These processes are prerequisites for exiting cargo form the ER. Later on, a fatty acid chain is attached to the cysteine located on the end of the transmembrane region and HEF is cleaved into 2 subunits, this process is essential for virus replication.[2]
In influenza viruses
In comparison, influenza A, B, and C have different spike proteins, the haemagglutinin and the neuraminidase. The HEF surface glycoprotein of influenza C consists of three activities, receptor-binding, receptor-inactivating, and fusion activity. Receptor-binding mediates the attachment of the virus to N-acetyl-9-O-acetylneuraminic acid on the cell surface, receptor-inactivating releases the 9-O-acetyl group from N-acetyl-9-O-acetylneuraminic acid and the fusion activity depends on the post-translational proteolytic cleavage of HEF into two subunits as well as exposure to an acidic environment. In low pH conditions, a conformational change of HEF occurs. In influenza A, the rearrangement of hydrophobic sequences at the N-terminus of subunit HEF2 becomes exposed and induces the fusion of the viral envelope with the membrane of the target cell.[9] Another way that fuses the viral envelope to the host cell is with endocytic vesicles. HEF does not cleave the terminal silica acid residue from carbohydrates but removes the acetyl group from the position C9 of N-acetyl-9-O-acetylneuraminic acid. This is required to release fresh budded virus particles from infected cells, which otherwise would be trapped in the plasma membrane if the receptor is still present [2]
Influenza C is distinguishable from influenza A and B by its structural components. There are three amino acids that comprises the cytoplasmic portion of HEF, arginine-threonine-lysine, whereas in influenza A and B consists of ten hemagglutinin amino acids. A post-translational modification of HEF is the acylation with fatty acids. The fatty acid, stearic acid, was detected to be the prevailing fatty acid attached to HEF, whereas the fatty acid palmitic acid was found in all other membrane proteins.[9] Due to the frequent reassortment of strains, it's monosubtypic and stable. This leads to a new strain that aids the virus in adapting better to its host.[2]
References
- 1 2 3 Zeng Q, Langereis MA, van Vliet AL, Huizinga EG, de Groot RJ (July 2008). "Structure of coronavirus hemagglutinin-esterase offers insight into corona and influenza virus evolution". Proceedings of the National Academy of Sciences of the United States of America. 105 (26): 9065–9. Bibcode:2008PNAS..105.9065Z. doi:10.1073/pnas.0800502105. PMC 2449365. PMID 18550812.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 Wang M, Ludwig K, Böttcher C, Veit M (May 2016). "The role of stearate attachment to the hemagglutinin-esterase-fusion glycoprotein HEF of influenza C virus". Cellular Microbiology. 18 (5): 692–704. doi:10.1111/cmi.12541. PMID 26518983.
- ↑ Yu J, Hika B, Liu R, Sheng Z, Hause BM, Li F, Wang D (July 2017). "The Hemagglutinin-Esterase Fusion Glycoprotein Is a Primary Determinant of the Exceptional Thermal and Acid Stability of Influenza D Virus". mSphere. 2 (4). doi:10.1128/mSphere.00254-17. PMC 5549178. PMID 28808690.
- 1 2 3 Langereis MA, Zeng Q, Gerwig GJ, Frey B, von Itzstein M, Kamerling JP, de Groot RJ, Huizinga EG (September 2009). "Structural basis for ligand and substrate recognition by torovirus hemagglutinin esterases". Proceedings of the National Academy of Sciences of the United States of America. 106 (37): 15897–902. Bibcode:2009PNAS..10615897L. doi:10.1073/pnas.0904266106. PMC 2747215. PMID 19721004.
- ↑ Copeland CS, Zimmer KP, Wagner KR, Healey GA, Mellman I, Helenius A (April 1988). "Folding, trimerization, and transport are sequential events in the biogenesis of influenza virus hemagglutinin". Cell. 53 (2): 197–209. doi:10.1016/0092-8674(88)90381-9. PMID 3359486. S2CID 46735853.
- 1 2 3 4 Sugahara K, Hongo S, Sugawara K, Li ZN, Tsuchiya E, Muraki Y, Matsuzaki Y, Nakamura K (June 2001). "Role of individual oligosaccharide chains in antigenic properties, intracellular transport, and biological activities of influenza C virus hemagglutinin-esterase protein". Virology. 285 (1): 153–64. doi:10.1006/viro.2001.0952. PMID 11414815.
- ↑ Martin LT, Verhagen A, Varki A (2003). "Recombinant influenza C hemagglutinin-esterase as a probe for sialic acid 9-O-acetylation". Recognition of Carbohydrates in Biological Systems, Part B: Specific Applications. Methods in Enzymology. Vol. 363. pp. 489–98. doi:10.1016/S0076-6879(03)01074-7. ISBN 9780121822668. PMID 14579598.
- 1 2 Wang M, Ludwig K, Böttcher C, Veit M (May 2016). "The role of stearate attachment to the hemagglutinin-esterase-fusion glycoprotein HEF of influenza C virus". Cellular Microbiology. 18 (5): 692–704. doi:10.1111/cmi.12541. PMID 26518983.
- 1 2 Szepanski S, Veit M, Pleschka S, Klenk HD, Schmidt MF, Herrler G (May 1994). "Post-translational folding of the influenza C virus glycoprotein HEF: defective processing in cells expressing the cloned gene". The Journal of General Virology. 75 (5): 1023–30. doi:10.1099/0022-1317-75-5-1023. PMID 8176364.