| Lipoxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Structure of rabbit reticulocyte 15S-lipoxygenase.[1] | |||||||||
| Identifiers | |||||||||
| Symbol | Lipoxygenase | ||||||||
| Pfam | PF00305 | ||||||||
| InterPro | IPR013819 | ||||||||
| PROSITE | PDOC00077 | ||||||||
| SCOP2 | 2sbl / SCOPe / SUPFAM | ||||||||
| OPM superfamily | 80 | ||||||||
| OPM protein | 2p0m | ||||||||
| |||||||||
Lipoxygenases (EC 1.13.11.-) (LOX) are a family of (non-heme) iron-containing enzymes most of which catalyze the dioxygenation of polyunsaturated fatty acids in lipids containing a cis,cis-1,4-pentadiene into cell signaling agents that serve diverse roles as autocrine signals that regulate the function of their parent cells, paracrine signals that regulate the function of nearby cells, and endocrine signals that regulate the function of distant cells.
The lipoxygenases are related to each other based upon their similar genetic structure and dioxygenation activity. However, one lipoxygenase, ALOXE3, while having a lipoxygenase genetic structure, possesses relatively little dioxygenation activity; rather its primary activity appears to be as an isomerase that catalyzes the conversion of hydroperoxy unsaturated fatty acids to their 1,5-epoxide, hydroxyl derivatives.
Lipoxygenases are found in eukaryotes (plants, fungi, animals, protists); while the third domain of terrestrial life, the archaea, possesses proteins with a slight (~20%) amino acid sequence similarity to lipoxygenases, these proteins lack iron-binding residues and therefore are not projected to possess lipoxygenase activity.[2]
Biochemistry
Based on detailed analyses of 15-lipoxygenase 1 and stabilized 5-lipoxygenase, lipoxygenase structures consist of a 15 kilodalton N-terminal beta barrel domain, a small (e.g. ~0.6 kilodalton) linker inter-domain (see Protein domain § Domains and protein flexibility), and a relatively large C-terminal catalytic domain which contains the non-heme iron critical for the enzymes' catalytic activity.[3] Most of the lipoxygenases (exception, ALOXE3) catalyze the reaction Polyunsaturated fatty acid + O2 → fatty acid hydroperoxide in four steps:
- the rate-limiting step of hydrogen abstraction from a bisallylic methylene carbon to form a fatty acid radical at that carbon
- rearrangement of the radical to another carbon center
- addition of molecular oxygen (O2) to the rearranged carbon radical center thereby forming a peroxy radical(—OO·) bond to that carbon
- reduction of the peroxy radical to its corresponding anion (—OO−)
The (—OO−) residue may then be protonated to form a hydroperoxide group (—OOH) and further metabolized by the lipoxygenase to e.g. leukotrienes, hepoxilins, and various specialized pro-resolving mediators, or reduced by ubiquitous cellular glutathione peroxidases to a hydroxy group thereby forming hydroxylated (—OH) polyunsaturated fatty acids such as the hydroxyeicosatetraenoic acids and HODEs (i.e. hydroxyoctadecaenoic acids).[3]
Polyunsaturated fatty acids that serve as substrates for one or more of the lipoxygenases include the omega 6 fatty acids, arachidonic acid, linoleic acid, dihomo-γ-linolenic acid, and adrenic acid; the omega-3 fatty acids, eicosapentaenoic acid, docosahexaenoic acid, and alpha-linolenic acid; and the omega-9 fatty acid, mead acid.[4] Certain types of the lipoxygenases, e.g. human and murine 15-lipoxygenase 1, 12-lipoxygenase B, and ALOXE3, are capable of metabolizing fatty acid substrates that are constituents of phospholipids, cholesterol esters, or complex lipids of the skin.[3] Most lipoxygenases catalyze the formation of initially formed hydroperoxy products that have S chirality. Exceptions to this rule include the 12R-lipoxygenases of humans and other mammals (see below).[3][4][5]
Lipoxygenases depend on the availability of their polyunsaturated fatty acid substrates which, particularly in mammalian cells, is normally maintained at extremely low levels. In general, various phospholipase A2s and diacylglycerol lipases are activated during cell stimulation, proceed to release these fatty acids from their storage sites, and thereby are key regulators in the formation of lipoxygenase-dependent metabolites.[3] In addition, cells, when so activated, may transfer their released polyunsaturated fatty acids to adjacent or nearby cells which then metabolize them through their lipoxygenase pathways in a process termed transcellular metabolism or transcellular biosynthesis.[6]
Biological function and classification
These enzymes are most common in plants where they may be involved in a number of diverse aspects of plant physiology including growth and development, pest resistance, and senescence or responses to wounding.[7] In mammals a number of lipoxygenases isozymes are involved in the metabolism of eicosanoids (such as prostaglandins, leukotrienes and nonclassic eicosanoids).[8] Sequence data is available for the following lipoxygenases:
Plant lipoxygenases
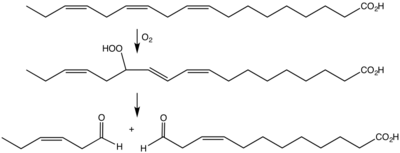
Plants express a variety of cytosolic lipoxygenases (EC 1.13.11.12; InterPro: IPR001246) as well as what seems to be a chloroplast isozyme.[9] Plant lipoxygenase in conjunction with hydroperoxide lyases are responsible for many fragrances and other signalling compounds. One example is cis-3-hexenal, the odor of freshly cut grass.
Human lipoxygenases
With the exception of the gene encoding 5-LOX (ALOX5), which is located on chromosome 10q11.2, all six human LOX genes are located on chromosome 17.p13 and code for a single chain protein of 75–81 kilodaltons that consists of 662–711 amino acids. Mammalian LOX genes contain 14 (ALOX5, ALOX12, ALOX15, ALOX15B) or 15 (ALOX12B, ALOXE3) exons with exon/intron boundaries at highly conserved positions.[11][12] The 6 human lipoxygenases along with some of the major products that they make, as well as some of their associations with genetic diseases, are as follows:[11][13][14][15][16]
- Arachidonate 5-lipoxygenase (ALOX5) (EC 1.13.11.34; InterPro: IPR001885), also termed 5-lipoxygenase, 5-LOX, and 5-LO. Major products: it metabolizes arachidonic acid to 5-hydroperoxy-eicostetraeoic acid (5-HpETE) which is converted to 1) 5-hydroxyicosatetraenoic acid (5-HETE) and then to 5-oxo-eicosatetraenoic acid (5-oxo-ETE), 2) leukotriene A4 (LTA4) which may then be converted to leukotriene B4 (LTB4) or leukotriene C4 (LTC4) (LTC4 may be further metabolized to leukotriene D4 [LTD4] and then to leukotriene E4 [LTE4]), or 3) acting in series with ALOX15, to the specialized pro-resolving mediators, lipoxins A4 and B4. ALOX5 also metabolizes eicosapentaenoic acid to a set of metabolites that contain 5 double bounds (i.e. 5-HEPE, 5-oxo-EPE, LTB5, LTC5, LTD5, and LTE5) as opposed to the 4 double bond-containing arachidonic acid metabolites. The enzyme, when acting in series with other lipoxygenase, cyclooxygenase, or cytochrome P450 enzymes, contributes to the metabolism of eicosapentaenoic acid to E series resolvins (see Resolvin § Resolvin Es) and of docosahexaenoic acid to D series resolvins (see Resolvin § Resolvin Ds). These resolvins are also classified as specialized pro-resolving mediators.
- Arachidonate 12-lipoxygenase (ALOX12) (EC 1.13.11.31; InterPro: IPR001885), also termed 12-lipoxygenase, platelet type platelet lipoxygenase (or 12-lipoxygenase, platelet type) 12-LOX, and 12-LO. It metabolizes arachidonic acid to 12-hydroperoxyeiocsatetraeoic acid (12-HpETE) which is further metabolized to 12-hydroxyeicosatetraenoic acid (12-HETE) or to various hepoxilins (also see 12-hydroxyeicosatetraenoic acid).
- Arachidonate 15-lipoxygenase-1 (ALOX15) (EC 1.13.11.33; InterPro: IPR001885), also termed 15-lipoxygenase-1, erythrocyte type 15-lipoxygenase (or 15-lipoxygenase, erythrocyte type), reticulocyte type 15-lipoxygenase (or 15-lipoxygenase, reticulocyte type), 15-LO-1, and 15-LOX-1. It metabolizes arachidonic acid principally to 1) 15-hydroperoxyeiocatetraenoic acid (15-HpETE) which is further metabolized to 15-hydroxyicosatetraenoic acid (15-HETE) but also to far smaller amounts of 2) 12-hydroperoxyeicosatetraenoic acid (12-HpETE) which is further metabolized to 12-hydroxyeicosatetraenoic acid and possibly the hepoxilins. ALOX15 actually prefers linoleic acid over arachidonic acid, metabolizing linoleic acid to 12-hydroperoxyoctadecaenoic acid (13-HpODE) which is further metabolized to 13-hydroxyoctadecadienoic acid (13-HODE). ALOX15 can metabolize polyunsaturated fatty acids that are esterified to phospholipids and/or to the cholesterol, i.e. cholesterol esters, in lipoproteins. This property along with its dual specificity in metabolizing arachidonic acid to 12-HpETE and 15-HpETE are similar to those of mouse Alox15 and has led to both enzymes being termed 12/15-lipoxygenases.
- Arachidonate 15-lipoxygenase type II (ALOX15B), also termed 15-lipoxygenase-2, 15-LOX-2, and 15-LOX-2.[17] It metabolizes arachidonic acid to 15-hydroperoxyeicosatetraenoic (15-HpETE) which is further metabolized to 15-hydroxyicosatetraenoic acid. ALOX15B has little or no ability to metabolize arachidonic acid to 12-hydroperoxeiocosatetraenoic acid (12-(HpETE) and only minimal ability to metabolize linoleic acid to 13-hydroperoxyoctadecaenoic acid (13-HpODE).
- Arachidonate 12-lipoxygenase, 12R type (ALOX12B), also termed 12R-lipoxygenase, 12R-LOX, and 12R-LO.[18] It metabolizes arachidonic acid to 12R-hydroxyeicosatetraenoic acid but does so only with low catalytic activity; its most physiologically important substrate is thought to be a sphingosine which contains a very long chain (16-34 carbons) omega-hydroxyl fatty acid that is in amide linkage to the sn-2 nitrogen of sphingosine at its carboxy end and esterified to linoleic acid at its omega hydroxyl end. In skin epidermal cells, ALOX12B metabolizes the linoleate in this esterified omega-hydroxyacyl-sphingosine (EOS) to its 9R-hydroperoxy analog. Inactivating mutations of ALOX12B are associated with the human skin disease, autosomal recessive congenital ichthyosiform erythroderma (ARCI).[18][19]
- Epidermis-type lipoxygenase (ALOXE3), also termed eLOX3 and lipoxygenase, epidermis type.[20] Unlike other lipoxygenases, ALOXE3 exhibits only a latent dioxygenase activity. Rather, its primary activity is as a hydroperoxide isomerase that metabolizes certain unsaturated hydroperoxy fatty acids to their corresponding epoxy alcohol and epoxy keto derivatives and thereby is also classified as a hepoxilin synthase. While it can metabolize 12S-hydroperoxyeicosatetraenoic acid (12S-HpETE) to the R stereoisomers of hepoxilins A3 and B3, ALOXE3 favors metabolizing R hydroperoxy unsaturated fatty acids and efficiently converts the 9(R)-hydroperoxy analog of EOS made by ALOX15B to its 9R(10R),13R-trans-epoxy-11E,13R and 9-keto-10E,12Z EOS analogs.[19] ALOXE3 is thought to act with ALOX12B in skin epidermis to form the latter two EOS analogs; inactivation mutations of ALOX3 are, similar to inactivating mutations in ALOX12B, associated with autosomal recessive congenital ichthyosiform erythroderma in humans.[19][20] Inactivating mutations in ALOX3 are also associated with the human disease lamellar ichthyosis (see Ichthyosis § Types – item 5 in the table).
Two lipoxygenases may act in series to make di-hydroxy or tri-hydroxy products that have activities quite different than either lipoxyenases' products. This serial metabolism may occur in different cell types that express only one of the two lipoxygenases in a process termed transcellular metabolism. For example, ALOX5 and ALOX15 or, alternatively, ALOX5 and ALOX12 can act serially to metabolize arachidonic acid into lipoxins (see 15-Hydroxyeicosatetraenoic acid §§ Further metabolism and Activities of 15(S)-HpETE, 15(S)-HETE, 15(R)-HpETE, 15(R)-HETE, and 15-oxo-ETE and Lipoxin § Synthesis) while ALOX15 and possibly ALOX15B can act with ALOX5 to metabolize eicosapentaenoic acid to resolvin D's (see Resolvin § Biochemistry and production).
Mouse lipoxygenases
The mouse is a common model to examine lipoxygenase function. However, there are some key differences between the lipoxygenases between mice and men that make extrapolations from mice studies to humans difficult. In contrast to the 6 functional lipoxygenases in humans, mice have 7 functional lipoxygenases and some of the latter have different metabolic activities than their human orthologs.[11][19][21] In particular, mouse Alox15, unlike human ALOX15, metabolizes arachidonic acid mainly to 12-HpETE and mouse Alox15b, in contrast to human ALOX15b, is primarily an 8-lipoxygenase, metabolizing arachdionic acid to 8-HpETE; there is no comparable 8-HpETE-forming lipoxygenase in humans.[22]
- Alox5 appears to be similar in function to human ALOX5.
- Alox12 differs from human ALOX12, which preferentially metabolizes arachidonic acid to 12-HpETE but also to substantial amounts of 15-HpETE, in that metabolizes arachidonic acid almost exclusively to 12-HpETE.
- Alox15 (also termed leukocyte-type 12-Lox, 12-Lox-l, and 12/15-Lox) differs from human ALOX15, which under standard assay conditions metabolizes arachidonic acid to 15-HpETE and 12-HpETE products in an 89 to 11 ratio, metabolizes arachidonic acid to 15-Hpete and 12-HpETE in a 1 to 6 ratio, i.e. its principal metabolite is 12-HpETE. Also, human ALOX15 prefers linoleic acid over arachidonic acid as a substrate, metabolizing it to 13-HpODE while Alox15 has little or no activity on linoleic acid. Alox15 can metabolize polyunsaturated fatty acids that are esterified to phospholipids and cholesterol (i.e. cholesterol esters). This property along with its dual specificity in metabolizing arachidonic acid to 12-HpETE and 15-HpETE are similar to those of human ALOX15 and has led to both enzymes being termed 12/15-lipoxygenases.
- Alox15b (also termed 8-lipoxygenase, 8-lox, and 15-lipoxygenase type II), in contrast to ALOX15B which metabolizes arachidonic acid principally to 15-HpETE and to a lesser extent linoleic acid to 13-HpODE, metabolizes arachidonic acid principally to 8S-HpETE and linoleic acid to 9-HpODE. Alox15b is as effective as ALOX5 in metabolizing 5-HpETE to leukotrienes.
- Alox12e (12-Lox-e, epidermal-type 12-Lox) is an ortholog to the human ALOX12P gene which has suffered damaging mutations and is not expressed. ALox12e prefers methyl esters over non-esterified polyunsaturated fatty acid substrates, metabolizing linoleic acid ester to its 13-hydroperoxy counterpart and to a lesser extent arachidonic acid ester to its 12-hydroperoxy counterpart.
- Alox12b (e-LOX2, epidermis-type Lox-12) appears to act similarly to ALOX12B to metabolize the linoleic acid moiety of EOS to its 9R-hydroperoxy counterpart and thereby contribute to skin integrity and water impermeability; mice depleted to Alox12b develop a severe skin defect similar to Congenital ichthyosiform erythroderma. Unlike human ALOX12B which cam metabolize arachidonic acid to 12R-HETE at a low rate, Alox12b does not metabolize arachidonic acid as free acid but dose metabolize arachidonic acid methyl ester to its 12R-hydroperoxy counterpart.
- Aloxe3 (epidermis-type Lox-3, eLox3) appears to act similarly to ALOXe3 in metabolizing the 9R-hydoperoxy-linoleate derivative of EOS to its epoxy and keto derivatives and to be involved in maintaining skin integrity and water impermeability. AloxE3 deletion leads to a defect similar to congenital ichthyosiform erythroderma.

3D structure
There are several lipoxygenase structures known including: soybean lipoxygenase L1 and L3, coral 8-lipoxygenase, human 5-lipoxygenase, rabbit 15-lipoxygenase and porcine leukocyte 12-lipoxygenase catalytic domain. The protein consists of a small N-terminal PLAT domain and a major C-terminal catalytic domain (see Pfam database), which contains the active site. In both plant and mammalian enzymes, the N-terminal domain contains an eight-stranded antiparallel β-barrel, but in the soybean lipoxygenases this domain is significantly larger than in the rabbit enzyme. The plant lipoxygenases can be enzymatically cleaved into two fragments which stay tightly associated while the enzyme remains active; separation of the two domains leads to loss of catalytic activity. The C-terminal (catalytic) domain consists of 18-22 helices and one (in rabbit enzyme) or two (in soybean enzymes) antiparallel β-sheets at the opposite end from the N-terminal β-barrel.
Active site
The iron atom in lipoxygenases is bound by four ligands, three of which are histidine residues.[23] Six histidines are conserved in all lipoxygenase sequences, five of them are found clustered in a stretch of 40 amino acids. This region contains two of the three zinc-ligands; the other histidines have been shown[24] to be important for the activity of lipoxygenases.
The two long central helices cross at the active site; both helices include internal stretches of π-helix that provide three histidine (His) ligands to the active site iron. Two cavities in the major domain of soybean lipoxygenase-1 (cavities I and II) extend from the surface to the active site. The funnel-shaped cavity I may function as a dioxygen channel; the long narrow cavity II is presumably a substrate pocket. The more compact mammalian enzyme contains only one boot-shaped cavity (cavity II). In soybean lipoxygenase-3 there is a third cavity which runs from the iron site to the interface of the β-barrel and catalytic domains. Cavity III, the iron site and cavity II form a continuous passage throughout the protein molecule.
The active site iron is coordinated by Nε of three conserved His residues and one oxygen of the C-terminal carboxyl group. In addition, in soybean enzymes the side chain oxygen of asparagine is weakly associated with the iron. In rabbit lipoxygenase, this Asn residue is replaced with His which coordinates the iron via Nδ atom. Thus, the coordination number of iron is either five or six, with a hydroxyl or water ligand to a hexacoordinate iron.
Details about the active site feature of lipoxygenase were revealed in the structure of porcine leukocyte 12-lipoxygenase catalytic domain complex[23][25] In the 3D structure, the substrate analog inhibitor occupied a U-shaped channel open adjacent to the iron site. This channel could accommodate arachidonic acid without much computation, defining the substrate binding details for the lipoxygenase reaction. In addition, a plausible access channel, which intercepts the substrate binding channel and extended to the protein surface could be counted for the oxygen path.
Biochemical classification
| EC 1.13.11.12 | lipoxygenase | (linoleate:oxygen 13-oxidoreductase) | linoleate + O2 = (9Z,11E,13S)-13-hydroperoxyoctadeca-9,11-dienoate |
| EC 1.13.11.31 | arachidonate 12-lipoxygenase | (arachidonate:oxygen 12-oxidoreductase) | arachidonate + O2 = (5Z,8Z,10E,12S,14Z)-12-hydroperoxyicosa-5,8,10,14-tetraenoate |
| EC 1.13.11.33 | arachidonate 15-lipoxygenase | (arachidonate:oxygen 15-oxidoreductase) | arachidonate + O2 = (5Z,8Z,11Z,13E,15S)-15-hydroperoxyicosa-5,8,11,13-tetraenoate |
| EC 1.13.11.34 | arachidonate 5-lipoxygenase | (arachidonate:oxygen 5-oxidoreductase) | arachidonate + O2 = leukotriene A4 + H2 |
| EC 1.13.11.40 | arachidonate 8-lipoxygenase | (arachidonate:oxygen 8-oxidoreductase) | arachidonate + O2 = (5Z,8R,9E,11Z,14Z)-8-hydroperoxyicosa-5,9,11,14-tetraenoate |
Soybean Lipoxygenase 1 exhibits the largest H/D kinetic isotope effect (KIE) on kcat (kH/kD) (81 near room temperature) so far reported for a biological system. Recently, an extremely elevated KIE of 540 to 730 was found in a double mutant Soybean Lipoxygenase 1.[26] Because of the large magnitude of the KIE, Soybean Lipoxygenase 1 has served as the prototype for enzyme-catalyzed hydrogen-tunneling reactions.
Human proteins expressed from the lipoxygenase family include ALOX12, ALOX12B, ALOX15, ALOX15B, ALOX5, and ALOXE3. While humans also possess the ALOX12P2 gene, which is an ortholog of the well-expressed Alox12P gene in mice, the human gene is a pseudogene; consequently, ALOX12P2 protein is not detected in humans.[27]
References
- ↑ Choi J, Chon JK, Kim S, Shin W (February 2008). "Conformational flexibility in mammalian 15S-lipoxygenase: Reinterpretation of the crystallographic data". Proteins. 70 (3): 1023–32. doi:10.1002/prot.21590. PMID 17847087. S2CID 40013415.
- ↑ Powell WS, Rokach J (2015). "Biosynthesis, biological effects, and receptors of hydroxyeicosatetraenoic acids (HETEs) and oxoeicosatetraenoic acids (oxo-ETEs) derived from arachidonic acid". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1851 (4): 340–55. doi:10.1016/j.bbalip.2014.10.008. PMC 5710736. PMID 25449650.
- 1 2 3 4 5 Kuhn H, Banthiya S, van Leyen K (2015). "Mammalian lipoxygenases and their biological relevance". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1851 (4): 308–30. doi:10.1016/j.bbalip.2014.10.002. PMC 4370320. PMID 25316652.
- 1 2 Gabbs M, Leng S, Devassy JG, Monirujjaman M, Aukema HM (2015). "Advances in Our Understanding of Oxylipins Derived from Dietary PUFAs". Advances in Nutrition. 6 (5): 513–40. doi:10.3945/an.114.007732. PMC 4561827. PMID 26374175.
- ↑ Mashima R, Okuyama T (2015). "The role of lipoxygenases in pathophysiology; new insights and future perspectives". Redox Biology. 6: 297–310. doi:10.1016/j.redox.2015.08.006. PMC 4556770. PMID 26298204.
- ↑ Capra V, Rovati GE, Mangano P, Buccellati C, Murphy RC, Sala A (2015). "Transcellular biosynthesis of eicosanoid lipid mediators". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1851 (4): 377–82. doi:10.1016/j.bbalip.2014.09.002. PMID 25218301.
- ↑ Vick BA, Zimmerman DC (1987). "Oxidative Systems for Modification of Fatty Acids: The Lipoxygenase Pathway". Oxidative systems for the modification of fatty acids: The Lipoxygenase Pathway. Vol. 9. pp. 53–90. doi:10.1016/b978-0-12-675409-4.50009-5. ISBN 9780126754094.
- ↑ Needleman P, Turk J, Jakschik BA, Morrison AR, Lefkowith JB (1986). "Arachidonic acid metabolism". Annu. Rev. Biochem. 55: 69–102. doi:10.1146/annurev.bi.55.070186.000441. PMID 3017195.
- ↑ Tanaka K, Ohta H, Peng YL, Shirano Y, Hibino T, Shibata D (1994). "A novel lipoxygenase from rice. Primary structure and specific expression upon incompatible infection with rice blast fungus". J. Biol. Chem. 269 (5): 3755–3761. doi:10.1016/S0021-9258(17)41924-7. PMID 7508918.
- ↑ KenjiMatsui (2006). "Green leaf volatiles: hydroperoxide lyase pathway of oxylipin metabolism". Current Opinion in Plant Biology. 9 (3): 274–280. doi:10.1016/j.pbi.2006.03.002. PMID 16595187.
- 1 2 3 Krieg, P; Fürstenberger, G (2014). "The role of lipoxygenases in epidermis". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1841 (3): 390–400. doi:10.1016/j.bbalip.2013.08.005. PMID 23954555.
- ↑ "ALOX5 arachidonate 5-lipoxygenase [Homo sapiens (human)] - Gene - NCBI".
- ↑ Haeggström, J. Z.; Funk, C. D. (2011). "Lipoxygenase and leukotriene pathways: Biochemistry, biology, and roles in disease". Chemical Reviews. 111 (10): 5866–98. doi:10.1021/cr200246d. PMID 21936577.
- ↑ Barden AE, Mas E, Mori TA (2016). "n-3 Fatty acid supplementation and proresolving mediators of inflammation". Current Opinion in Lipidology. 27 (1): 26–32. doi:10.1097/MOL.0000000000000262. PMID 26655290. S2CID 45820130.
- ↑ Qu Q, Xuan W, Fan GH (2015). "Roles of resolvins in the resolution of acute inflammation". Cell Biology International. 39 (1): 3–22. doi:10.1002/cbin.10345. PMID 25052386. S2CID 10160642.
- ↑ Romano M, Cianci E, Simiele F, Recchiuti A (2015). "Lipoxins and aspirin-triggered lipoxins in resolution of inflammation". European Journal of Pharmacology. 760: 49–63. doi:10.1016/j.ejphar.2015.03.083. PMID 25895638.
- ↑ "WikiGenes - Collaborative Publishing". WikiGenes - Collaborative Publishing. Retrieved 17 April 2018.
- 1 2 "WikiGenes - Collaborative Publishing". WikiGenes - Collaborative Publishing. Retrieved 17 April 2018.
- 1 2 3 4 Muñoz-Garcia, A; Thomas, C. P.; Keeney, D. S.; Zheng, Y; Brash, A. R. (2014). "The importance of the lipoxygenase-hepoxilin pathway in the mammalian epidermal barrier". Biochimica et Biophysica Acta (BBA) - Molecular and Cell Biology of Lipids. 1841 (3): 401–8. doi:10.1016/j.bbalip.2013.08.020. PMC 4116325. PMID 24021977.
- 1 2 "WikiGenes - Collaborative Publishing". WikiGenes - Collaborative Publishing. Retrieved 17 April 2018.
- ↑ Taylor, P. R.; Heydeck, D; Jones, G. W.; Krönke, G; Funk, C. D.; Knapper, S; Adams, D; Kühn, H; O'Donnell, V. B. (2012). "Development of myeloproliferative disease in 12/15-lipoxygenase deficiency". Blood. 119 (25): 6173–4, author reply 6174–5. doi:10.1182/blood-2012-02-410928. PMC 3392071. PMID 22730527.
- ↑ Cole, B. K.; Lieb, D. C.; Dobrian, A. D.; Nadler, J. L. (2013). "12- and 15-lipoxygenases in adipose tissue inflammation". Prostaglandins & Other Lipid Mediators. 104–105: 84–92. doi:10.1016/j.prostaglandins.2012.07.004. PMC 3526691. PMID 22951339.
- 1 2 Boyington JC, Gaffney BJ, Amzel LM (1993). "The three-dimensional structure of an arachidonic acid 15-lipoxygenase". Science. 260 (5113): 1482–1486. Bibcode:1993Sci...260.1482B. doi:10.1126/science.8502991. PMID 8502991.
- ↑ Steczko J, Donoho GP, Clemens JC, Dixon JE, Axelrod B (1992). "Conserved histidine residues in soybean lipoxygenase: functional consequences of their replacement". Biochemistry. 31 (16): 4053–4057. doi:10.1021/bi00131a022. PMID 1567851.
- ↑ Xu, S.; Mueser T.C.; Marnett L.J.; Funk M.O. (2012). "Crystal structure of 12-lipoxygenase catalytic-domain-inhibitor complex identifies a substrate-binding channel for catalysis". Structure. 20 (9): 1490–7. doi:10.1016/j.str.2012.06.003. PMC 5226221. PMID 22795085.
- ↑ Hu, S; Sharma, S. C.; Scouras, A. D.; Soudackov, A. V.; Carr, C. A.; Hammes-Schiffer, S; Alber, T; Klinman, J. P. (2014). "Extremely elevated room-temperature kinetic isotope effects quantify the critical role of barrier width in enzymatic C-H activation". Journal of the American Chemical Society. 136 (23): 8157–60. doi:10.1021/ja502726s. PMC 4188422. PMID 24884374.
- ↑ "WikiGenes - Collaborative Publishing". WikiGenes - Collaborative Publishing. Retrieved 17 April 2018.
External links
- LOX-DB – LipOXygenases DataBase
- Lipoxygenases iron-binding region in PROSITE
- PDB: 1YGE – structure of lipoxygenase-1 from soybean (Glycine max)
- PDB: 1IK3 – structure of soybean lipoxygenase-3 in complex with (9Z,11E,13S)-13-hydroperoxyoctadeca-9,11-dienoic acid
- PDB: 1LOX – structure of rabbit 15-lipoxygenase in complex with inhibitor
- PDB: 3RDE – structure of the catalytic domain of porcine leukocyte 12-lipoxygenasean with inhibitor
- UMich Orientation of Proteins in Membranes families/superfamily-87 – animal lipoxygenases
- Lipoxygenase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Blanch Time and Cultivar Effects on Quality of Frozen and Stored Corn and Broccoli – lipoxygenase, peroxidase, cystine lyase enzyme inactivation in blanching