Tc1/mariner is a class and superfamily of interspersed repeats DNA (Class II) transposons.[1] The elements of this class are found in all animals,[2] including humans. They can also be found in protists and bacteria.[3][4]
The class is named after its two best-studied members, the Tc1 transposon of Caenorhabditis elegans and the mariner transposon of Drosophila.
Structure
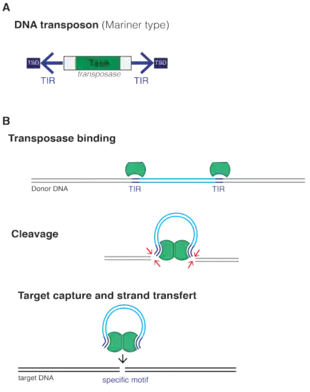
The transposon consists of a transposase gene, flanked by two terminal inverted repeats (TIR). Two short tandem site duplications (TSD) are present on both sides of the insert. Transposition happens when two transposases recognize and bind to TIR sequences, join together and promote DNA double-strand cleavage. The DNA-transposase complex then inserts its DNA cargo at specific DNA motifs elsewhere in the genome, creating short TSDs upon integration.[5] In the IS630/Tc1/mariner system, the motif used is a "TA" dinucleotide, duplicated on both ends after insertion.
When the transposase gene is not carried by the transposon, it becomes a non-autonomous in that it now requires the gene to be expressed elsewhere to move around.
Transposase
| Transposase, type 1 (partial DDE domain) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Transpotase_1 | ||||||||
| Pfam | PF01359 | ||||||||
| Pfam clan | CL0219 | ||||||||
| InterPro | IPR001888 | ||||||||
| CATH | 4u7b | ||||||||
| |||||||||
| HTH domain in Mos1 transposase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | HTH_48 | ||||||||
| Pfam | PF17906 | ||||||||
| Pfam clan | CL0123 | ||||||||
| InterPro | IPR041426 | ||||||||
| CATH | 4u7b | ||||||||
| |||||||||

The 360-amino acid polypeptide has three major subdomains: the amino-terminal DNA-recognition domain that is responsible for binding to the DR sequences in the mirrored IR/DR sequences of the transposon, a nuclear localization sequence (NLS), and a DDE domain that catalyzes the cut-and-paste set of reactions that comprise transposition. The DNA-recognition domain has two paired box sequences that can bind to DNA and are related to various motifs found on some transcription factors; the two paired boxes are labeled PAI and RED, both having the helix-turn-helix motif common for DNA-binding domains. The catalytic domain has the hallmark DDE (sometimes DDD) amino acids that are found in many transposase and recombinase enzymes. In addition, there is a region that is highly enriched in glycine (G) amino acids.
Several signatures for the superfamily of transcriptases have been given in various domain databases given the multi-domain nature of the protein. In addition, each domain are often represented by multiple entries, such as PF17906/PF01710/PF11427 among others for the "PAI" half of the box. The RED box is similarly diverse (PF08279/PF13412/PF01498, etc.) and is often in a winged HTH form for DNA recognition.
Sub-groups
The Tc1/mariner superfamily is generally subdivided by the catalytic domains of its transposase. It generally use a DDE (Asp-Asp-Glu) or DDD catalytic triad.
Tc1
Tc1 (DD34E) is a transposon active in Caenorhabditis elegans.[6][7] There are also Tc1-like transposons in humans, all inactive. Tc1-like elements are present in other lower vertebrates, including several fish species and amphibians.[8]
In C. elegans, it is a 1610 base-pair long sequence.[9] Experiments show that this element "jumps" in human cells, with its transposase as the only protein required.[10]
Another example of this family is Tc3, also a transposon found in C. elegans.[11]
Mariner
Mariner (DD34D) elements are found in multiple species, including humans.[12][13] The Mariner transposon was first discovered by Jacobson and Hartl in Drosophila in 1986.[14] A classification of the group was published in 1993, which divided such sequences in insects into the mauritiana, cecropia, mellifera, irritans, and capitata subfamilies, after the types of insects they are found in.[15] The classification does extend to other species.
This transposable element is known for its uncanny ability to be transmitted horizontally in many species.[16][17] There are an estimated 14,000 copies of Mariner in the human genome comprising 2.6 million base pairs.[18] The first mariner-element transposons outside of animals were found in Trichomonas vaginalis.[19]
Human Mariner-like transposons are divided into Hsmar1 (cecropia) and Hsmar2 (irritans) subfamilies. Although both types are inactive, one copy of Hsmar1 found in the SETMAR gene is under selection as it provides DNA-binding for the histone-modifying protein.[20] Hsmar2 has been reconstructed multiple times from the fossil sequences.[21]
Mos1 (for Mosaic element) was discovered in Drosophila mauritiana.[22] The Himar1 element has been isolated from the horn fly, Haematobia irritans and can be used as a genetic tool in Escherichia coli.[23]
Other families
The rosa (DD41D) family is a family found in Ceratitis rosa.[24] Pogo/Fot1 (DDxD) is yet another family in this superfamily, x indicating a variable length. IS630, a mobile element in Shigella sonnei, also belongs to this superfamily.[4]
A few additional new families with different lengths between the triads have been reported.[25]
Pogo, also known as Tigger in humans,[26] has been domesticated by humans and yeast alike into the CENPB gene. Other human domestications of pogo include TIGD1, TIGD2, TIGD3, TIGD4, TIGD5, TIGD6, TIGD7, JRK, JRKL, POGK, and POGZ.[27]
See also
References
- ↑ Plasterk, Ronald H.A; Izsvák, Zsuzsanna; Ivics, Zoltán (1999). "Resident aliens: The Tc1/mariner superfamily of transposable elements". Trends in Genetics. 15 (8): 326–32. doi:10.1016/S0168-9525(99)01777-1. PMID 10431195.
- ↑ Robertson, H.M. (1995). "The Tc1-mariner superfamily of transposons in animals". J. Insect Physiol. 41 (2): 99–105. doi:10.1016/0022-1910(94)00082-r.
- ↑ Bradic, Martina; Warring, Sally D; Low, Vivien; Carlton, Jane M (2014). "The Tc1/mariner transposable element family shapes genetic variation and gene expression in the protist Trichomonas vaginalis". Mobile DNA. 5: 12. doi:10.1186/1759-8753-5-12. PMC 4021607. PMID 24834134.
- 1 2 Capy, Pierre; Langin, Thierry; Higuet, Dominique; Maurer, Patricia; Bazin, Claude (1997). "Do the integrases of LTR-retrotransposons and class II element transposases have a common ancestor?". Genetica. 100 (1/3): 63–72. doi:10.1023/A:1018300721953. PMID 9440259. S2CID 24866580.
- 1 2 Walter, Marius (2016). Transposon regulation upon dynamic loss of DNA methylation (Thesis). Université Pierre et Marie Curie. doi:10.13140/rg.2.2.18747.21286.
- ↑ Babity, J. M.; Starr, T. V.; Rose, A. M. (1990). "Tc1 transposition and mutator activity in a Bristol strain of Caenorhabditis elegans". Molecular & General Genetics. 222 (1): 65–70. doi:10.1007/bf00283024. PMID 1978238. S2CID 11275388.
- ↑ Harris, L. J.; Rose, A. M. (1989). "Structural analysis of Tc1 elements in Caenorhabditis elegans var. Bristol (strain N2)". Plasmid. 22 (1): 10–21. doi:10.1016/0147-619X(89)90031-0. PMID 2550981.
- ↑ Goodier, John L.; Davidson, William S. (1994). "Tc1 Transposon-like Sequences are Widely Distributed in Salmonids". Journal of Molecular Biology. 241 (1): 26–34. doi:10.1006/jmbi.1994.1470. PMID 8051704.
- ↑ Rosenzweig, B; Liao, L. W.; Hirsh, D (1983). "Sequence of the C. Elegans transposable element Tc1". Nucleic Acids Research. 11 (12): 4201–4209. doi:10.1093/nar/11.12.4201. PMC 326035. PMID 6306578.
- ↑ Schouten, G. J.; Van Luenen, H. G. A. M.; Verra, N. C. V.; Valerio, D.; Plasterk, R. H. A. (1998). "Transposon Tc1 of the nematode Caenorhabditis elegans jumps in human cells". Nucleic Acids Research. 26 (12): 3013–7. doi:10.1093/nar/26.12.3013. PMC 147650. PMID 9611249.
- ↑ Vanluenen, H; Colloms, S; Plasterk, R (1994). "The mechanism of transposition of Tc3 in C. Elegans". Cell. 79 (2): 293–301. doi:10.1016/0092-8674(94)90198-8. PMID 7954797. S2CID 5804504.
- ↑ Oosumi, T; Belknap, WR; Garlick, B (14 December 1995). "Mariner transposons in humans". Nature. 378 (6558): 672. Bibcode:1995Natur.378..672O. doi:10.1038/378672a0. PMID 7501013. S2CID 4269186.
- ↑ Reiter, L. T.; Liehr, T; Rautenstrauss, B; Robertson, H. M.; Lupski, J. R. (1999). "Localization of mariner DNA Transposons in the Human Genome by PRINS". Genome Research. 9 (9): 839–843. doi:10.1101/gr.9.9.839. PMC 310809. PMID 10508842.
- ↑ Jacobson JW, Medhora MM, Hartl DL (November 1986). "Molecular structure of a somatically unstable transposable element in Drosophila". Proc. Natl. Acad. Sci. U.S.A. 83 (22): 8684–8. Bibcode:1986PNAS...83.8684J. doi:10.1073/pnas.83.22.8684. PMC 386995. PMID 3022302.
- ↑ Robertson, H. M.; MacLeod, E. G. (1993). "Five major subfamilies of mariner transposable elements in insects, including the Mediterranean fruit fly, and related arthropods". Insect Molecular Biology. 2 (3): 125–39. doi:10.1111/j.1365-2583.1993.tb00132.x. PMID 9087550. S2CID 11093292.
- ↑ Lohe AR, Moriyama EN, Lidholm DA, Hartl DL (January 1995). "Horizontal transmission, vertical inactivation, and stochastic loss of mariner-like transposable elements". Mol. Biol. Evol. 12 (1): 62–72. doi:10.1093/oxfordjournals.molbev.a040191. PMID 7877497.
- ↑ Lampe DJ, Witherspoon DJ, Soto-Adames FN, Robertson HM (April 2003). "Recent horizontal transfer of mellifera subfamily mariner transposons into insect lineages representing four different orders shows that selection acts only during horizontal transfer". Mol. Biol. Evol. 20 (4): 554–62. doi:10.1093/molbev/msg069. PMID 12654937.
- ↑ Mandal PK, Kazazian HH (October 2008). "SnapShot: Vertebrate transposons". Cell. 135 (1): 192–192.e1. doi:10.1016/j.cell.2008.09.028. PMID 18854165. S2CID 82147.
- ↑ Carlton JM, Hirt RP, Silva JC, et al. (January 2007). "Draft genome sequence of the sexually transmitted pathogen Trichomonas vaginalis". Science. 315 (5809): 207–12. Bibcode:2007Sci...315..207C. doi:10.1126/science.1132894. PMC 2080659. PMID 17218520.
- ↑ Miskey, C.; Papp, B.; Mates, L.; Sinzelle, L.; Keller, H.; Izsvak, Z.; Ivics, Z. (2 April 2007). "The Ancient mariner Sails Again: Transposition of the Human Hsmar1 Element by a Reconstructed Transposase and Activities of the SETMAR Protein on Transposon Ends". Molecular and Cellular Biology. 27 (12): 4589–4600. doi:10.1128/MCB.02027-06. PMC 1900042. PMID 17403897.
- ↑ Gil, Estel; Bosch, Assumpcio; Lampe, David; Lizcano, Jose M.; Perales, Jose C.; Danos, Olivier; Chillon, Miguel; Cordaux, Richard (11 September 2013). "Functional Characterization of the Human Mariner Transposon Hsmar2" (PDF). PLOS ONE. 8 (9): e73227. Bibcode:2013PLoSO...873227G. doi:10.1371/journal.pone.0073227. PMC 3770610. PMID 24039890.
- ↑ Hartl, D (2001). "Discovery of the transposable element mariner". Genetics. 157 (2): 471–476. doi:10.1093/genetics/157.2.471. PMC 1461507. PMID 11156971.
- ↑ Lampe, D. J.; Akerley, B. J.; Rubin, E. J.; Mekalanos, J. J.; Robertson, H. M. (1999). "Hyperactive transposase mutants of the Himar1 mariner transposon". Proceedings of the National Academy of Sciences. 96 (20): 11428–11433. Bibcode:1999PNAS...9611428L. doi:10.1073/pnas.96.20.11428. PMC 18050. PMID 10500193.
- ↑ Gomulski, LM; Torti, C; Bonizzoni, M; Moralli, D; Raimondi, E; Capy, P; Gasperi, G; Malacrida, AR (December 2001). "A new basal subfamily of mariner elements in Ceratitis rosa and other tephritid flies". Journal of Molecular Evolution. 53 (6): 597–606. Bibcode:2001JMolE..53..597G. doi:10.1007/s002390010246. PMID 11677619. S2CID 21289272.
- ↑ Shao, H; Tu, Z (November 2001). "Expanding the diversity of the IS630-Tc1-mariner superfamily: discovery of a unique DD37E transposon and reclassification of the DD37D and DD39D transposons". Genetics. 159 (3): 1103–15. doi:10.1093/genetics/159.3.1103. PMC 1461862. PMID 11729156.
- ↑ Smit, AF; Riggs, AD (20 February 1996). "Tiggers and DNA transposon fossils in the human genome". Proceedings of the National Academy of Sciences of the United States of America. 93 (4): 1443–8. doi:10.1073/pnas.93.4.1443. PMC 39958. PMID 8643651.
- ↑ Casola, C; Hucks, D; Feschotte, C (January 2008). "Convergent domestication of pogo-like transposases into centromere-binding proteins in fission yeast and mammals". Molecular Biology and Evolution. 25 (1): 29–41. doi:10.1093/molbev/msm221. PMC 2268608. PMID 17940212.