| oxalate decarboxylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
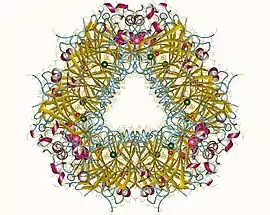 Oxalate decarboxylase hexamer, Bacillus subtilis | |||||||||
| Identifiers | |||||||||
| EC no. | 4.1.1.2 | ||||||||
| CAS no. | 9024-97-9 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
In enzymology, an oxalate decarboxylase (EC 4.1.1.2) is an oxalate degrading enzyme that catalyzes the chemical reaction
- oxalate + H+ formate + CO2
Thus, the two substrates of this enzyme are oxalate and H+, whereas its two products are formate and CO2.
This enzyme belongs to the family of lyases, specifically the carboxy-lyases, which cleave carbon-carbon bonds. The systematic name of this enzyme class is oxalate carboxy-lyase (formate-forming). This enzyme is also called oxalate carboxy-lyase. This enzyme participates in glyoxylate and dicarboxylate metabolism.
Structural studies
As of late 2007, 5 structures have been solved for this class of enzymes, with PDB accession codes 1UW8, 2UY8, 2UY9, 2UYA, and 2UYB.
References
- HAYAISHI O, JAKOBY WB, OHMURA E (1956). "Enzymatic decarboxylation of oxalic acid". J. Biol. Chem. 222 (1): 435–46. PMID 13367015.
- Tanner A, Bornemann S (2000). "Bacillus subtilis YvrK is an acid-induced oxalate decarboxylase". J. Bacteriol. 182 (18): 5271–3. doi:10.1128/JB.182.18.5271-5273.2000. PMC 94680. PMID 10960116.
- Tanner A, Bowater L, Fairhurst SA, Bornemann S (2001). "Oxalate decarboxylase requires manganese and dioxygen for activity Overexpression and characterization of Bacillus subtilis YvrK and YoaN". J. Biol. Chem. 276 (47): 43627–34. doi:10.1074/jbc.M107202200. PMID 11546787.
This article is issued from Wikipedia. The text is licensed under Creative Commons - Attribution - Sharealike. Additional terms may apply for the media files.