| pyruvate, phosphate dikinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 2.7.9.1 | ||||||||
| CAS no. | 9027-40-1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Pyruvate, phosphate dikinase | |
|---|---|
 | |
| Identifiers | |
| Symbol | PPDK |
| InterPro | IPR010121 |
| See domains below. | |
Pyruvate, phosphate dikinase, or PPDK (EC 2.7.9.1) is an enzyme in the family of transferases that catalyzes the chemical reaction
- ATP + pyruvate + phosphate AMP + phosphoenolpyruvate + diphosphate
This enzyme has been studied primarily in plants, but it has been studied in some bacteria as well.[1] It is a key enzyme in gluconeogenesis and photosynthesis that is responsible for reversing the reaction performed by pyruvate kinase in Embden-Meyerhof-Parnas glycolysis. It should not be confused with pyruvate, water dikinase.
It belongs to the family of transferases, to be specific, those transferring phosphorus-containing groups (phosphotransferases) with paired acceptors (dikinases). This enzyme participates in pyruvate metabolism and carbon fixation.
Nomenclature
The systematic name of this enzyme class is ATP:pyruvate, phosphate phosphotransferase. Other names in common use include pyruvate, orthophosphate dikinase, pyruvate-phosphate dikinase (phosphorylating), pyruvate phosphate dikinase, pyruvate-inorganic phosphate dikinase, pyruvate-phosphate dikinase, pyruvate-phosphate ligase, pyruvic-phosphate dikinase, pyruvic-phosphate ligase, pyruvate, Pi dikinase, and PPDK.
Reaction mechanism
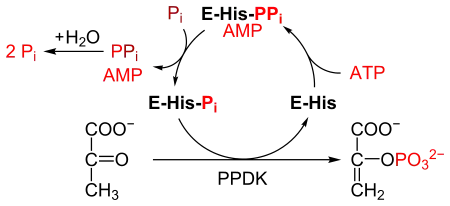
PPDK catalyses the conversion of pyruvate to phosphoenolpyruvate (PEP), consuming 1 molecule of ATP, and producing one molecule of AMP in the process. The mechanism consists of 3 reversible reactions:[2]
- The enzyme PPDK binds to ATP, to produce AMP and a diphosphorylated PPDK.
- The diphosphorylated PPDK binds to inorganic phosphate, producing diphosphate and (mono)phosphorylated PPDK.
- Phosphorylated PPDK binds to pyruvate, producing phosphoenolpyruvate, and regenerating PPDK.
The reaction is similar to the reaction catalysed by pyruvate kinase, which also converts pyruvate to PEP.[3] However, pyruvate kinase catalyses an irreversible reaction, and does not consume ATP. By contrast, PPDK catalyses a reversible reaction, and consumes 1 molecule of ATP for each molecule of pyruvate converted.
Currently, the details of each mechanistic step is unknown[3]
Structure
In its active form, PPDK is a homotetramer with subunits about 95 kDa[4]
There are two different reaction centres about 45 Angstroms apart, in which different substrates bind.[5] The nucleotide (ATP) binding site is on the N-terminus, has 240 amino acids, and a characteristic ATP-grasp. The pyruvate/PEP binding site is on the C-terminus, has 340 amino acids, and an α/β-barrel fold. There is also a central domain, which contains His455, the primary residue responsible for catalysis. His455 is the phosphoryl acceptor or donor residue.[3] The structure of the enzyme suggests that the His455 arm undergoes a swivelling motion to shuttle a phosphoryl group between the two reaction centres.[6] During this swivelling, the central domain rotates at least 92 degrees, and translates 0.5 Angstroms.[7]
Studies of crystal structures of PPDK show that the central domain is located in different proximity to the two other domains depending on the source of the enzyme.[7] In maize, it is closer to the C-terminal, while in Clostridium symbiosum, it is closer to the N-terminal.
Research has shown that the PPDK binding mechanisms are similar to that of D-Ala-D-Ala ligase and pyruvate kinase.[5] In particular, PPDK is very similar to pyruvate kinase, which also catalyses the conversion of pyruvate to phosphoenolpyruvate; however, it does so without a phosphorylated-enzyme intermediate.[3] Though their amino acid sequences are different, residues key to catalysis are preserved in both enzymes. Point-mutagenesis experiments have shown that catalytic residues include Arg561, Arg617, Glu745, Asn768, and Cys831 (numbering relative to the C, symbiosum protein, PDB: 1KBL, 1KC7).[3]
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Biological function and evolution
PPDK is used in the C4 pathway, to improve the efficiency of carbon dioxide fixation.[8] In environments where there is a lot of light, the rate of photosynthesis in plants is limited by the rate of carbon dioxide (CO2) uptake. This can be improved by using a series of chemical reactions to transport CO2 from mesophyll cells (which are located on the outside of a leaf) to bundle sheath cells (which are located inside the cells). PPDK converts pyruvate to PEP, which reacts with CO2 to produce oxaloacetate. When CO2 is released in the bundle sheath cells, pyruvate is regenerated, and the cycle continues.[8]
Though the reaction catalysed by PPDK is reversible, PEP is favoured as the product in biological conditions. This is due to the basic pH in the stroma, where the reaction occurs, as well as high concentrations of adenylate kinase and pyrophosphatase. Because these two enzymes catalyse exergonic reactions involving AMP, and disphosphate, respectively, they drive the PPDK-catalysed reaction forward.[9] Because PPDK consumes ATP, the C4 pathway is unfavourable for plants in environments with little access to light, as they are unable to produce large quantities of ATP.[8]
PPDK is highly abundant in C4 leaves, comprising up to 10% of total protein.[10] Research has shown that the enzyme is about 96% identical in different species of plants. Hybridization experiments revealed that the genetic differences correlate with the extent to which the plants perform the C4 pathway – the uncommon sequences exist in plants which also display C3 characteristics.[11] PPDK is also found in small quantities in C3 plants. Evolutionary history suggests that it once had a role in glycolysis like the similar pyruvate kinase, and eventually evolved into the C4 pathway.[10]
Besides plants, PPDK is also found in the parasitic ameoba Entamoeba histolytica (P37213) and the bacteria Clostridium symbiosum (P22983; as well as other bacteria).[12] In those two organisms PPDK functions similarly to (and sometimes in place of) pyruvate kinase, catalyzing the reaction in the ATP-producing direction as a part of glycolysis. Inhibitors for the Entamoeba PPDK have been proposed as amebicides against this organism.[13]
Regulation

Plant PPDK is regulated by the pyruvate, phosphate dikinase regulatory protein (PDRP).[4] When levels of light are high, PDRP dephosphorylates Thr456 on PPDK using AMP, thus activating the enzyme.[10] PDRP deactivates PPDK by phosphorylating the same threonine residue, using diphosphate. PDRP is a unique regulator because it catalyses both activation and deactivation of PPDK, through two different mechanisms.[10]
Research on maize PPDK suggests that introns, terminator sequences, and perhaps other enhancer sequences, act cooperatively to increase the level of functional and stable mRNA. PPDK cDNA was expressed only slightly in transgenic rice, compared to intact DNA which saw significant expression.[14]
Structural studies
As of early 2018, 14 structures have been solved for this class of enzymes, with PDB accession codes 1DIK, 1GGO, 1H6Z, 1JDE, 1KBL, 1KC7, 1VBG, 1VBH, 2DIK, 2FM4, 5JVJ, 5JVL, 5JVN, 5LU4.
References
- ↑ Pocalyko DJ, Carroll LJ, Martin BM, Babbitt PC, Dunaway-Mariano D (December 1990). "Analysis of sequence homologies in plant and bacterial pyruvate phosphate dikinase, enzyme I of the bacterial phosphoenolpyruvate: sugar phosphotransferase system and other PEP-utilizing enzymes. Identification of potential catalytic and regulatory motifs". Biochemistry. 29 (48): 10757–65. doi:10.1021/bi00500a006. PMID 2176881.
- ↑ Evans HJ, Wood HG (December 1968). "The mechanism of the pyruvate, phosphate dikinase reaction". Proceedings of the National Academy of Sciences of the United States of America. 61 (4): 1448–53. Bibcode:1968PNAS...61.1448E. doi:10.1073/pnas.61.4.1448. PMC 225276. PMID 4303480.
- 1 2 3 4 5 Herzberg O, Chen CC, Liu S, Tempczyk A, Howard A, Wei M, et al. (January 2002). "Pyruvate site of pyruvate phosphate dikinase: crystal structure of the enzyme-phosphonopyruvate complex, and mutant analysis". Biochemistry. 41 (3): 780–7. doi:10.1021/bi011799+. PMID 11790099.
- 1 2 Chastain CJ, Failing CJ, Manandhar L, Zimmerman MA, Lakner MM, Nguyen TH (May 2011). "Functional evolution of C(4) pyruvate, orthophosphate dikinase". Journal of Experimental Botany. 62 (9): 3083–91. doi:10.1093/jxb/err058. PMID 21414960.
- 1 2 Herzberg O, Chen CC, Kapadia G, McGuire M, Carroll LJ, Noh SJ, Dunaway-Mariano D (April 1996). "Swiveling-domain mechanism for enzymatic phosphotransfer between remote reaction sites". Proceedings of the National Academy of Sciences of the United States of America. 93 (7): 2652–7. Bibcode:1996PNAS...93.2652H. doi:10.1073/pnas.93.7.2652. PMC 39685. PMID 8610096.
- ↑ Lim K, Read RJ, Chen CC, Tempczyk A, Wei M, Ye D, et al. (December 2007). "Swiveling domain mechanism in pyruvate phosphate dikinase". Biochemistry. 46 (51): 14845–53. CiteSeerX 10.1.1.421.2653. doi:10.1021/bi701848w. PMID 18052212.
- 1 2 Nakanishi T, Nakatsu T, Matsuoka M, Sakata K, Kato H (February 2005). "Crystal structures of pyruvate phosphate dikinase from maize revealed an alternative conformation in the swiveling-domain motion". Biochemistry. 44 (4): 1136–44. doi:10.1021/bi0484522. PMID 15667207.
- 1 2 3 Berg J, Tymoczko J, Stryer L (2012). "The Calvin Cycle and the Pentose Phosphate Pathway". Biochemistry (7th ed.). New York: W.H Freeman. pp. 599–600. ISBN 9780716787242.
- ↑ Chastain C (2010). "Structure, Function, and Post-Translational Regulation of C4 Pyruvate Orthophosphate Dikinase". In Raghavendra A (ed.). C4 Photosynthesis and Related CO2 Concentrating Mechanisms. pp. 301–305. ISBN 9789048194063.
- 1 2 3 4 Chastain CJ, Fries JP, Vogel JA, Randklev CL, Vossen AP, Dittmer SK, et al. (April 2002). "Pyruvate,orthophosphate dikinase in leaves and chloroplasts of C(3) plants undergoes light-/dark-induced reversible phosphorylation". Plant Physiology. 128 (4): 1368–78. doi:10.1104/pp.010806. PMC 154264. PMID 11950985.
- ↑ Rosche E, Streubel M, Westhoff P (October 1994). "Primary structure of the photosynthetic pyruvate orthophosphate dikinase of the C3 plant Flaveria pringlei and expression analysis of pyruvate orthophosphate dikinase sequences in C3, C3-C4 and C4 Flaveria species". Plant Molecular Biology. 26 (2): 763–9. doi:10.1007/bf00013761. PMID 7948930. S2CID 23276817.
- ↑ UniProt 50%-90% clusters: From Clostridium PPDK
- ↑ Stephen P, Vijayan R, Bhat A, Subbarao N, Bamezai RN (September 2008). "Molecular modeling on pyruvate phosphate dikinase of Entamoeba histolytica and in silico virtual screening for novel inhibitors". Journal of Computer-Aided Molecular Design. 22 (9): 647–60. Bibcode:2008JCAMD..22..647S. doi:10.1007/s10822-007-9130-2. PMID 17710553. S2CID 25026913.
- ↑ Fukayama H, Tsuchida H, Agarie S, Nomura M, Onodera H, Ono K, et al. (November 2001). "Significant accumulation of C(4)-specific pyruvate, orthophosphate dikinase in a C(3) plant, rice". Plant Physiology. 127 (3): 1136–46. doi:10.1104/pp.010641. PMC 129282. PMID 11706193.
Further reading
- Hatch MD, Slack CR (January 1968). "A new enzyme for the interconversion of pyruvate and phosphopyruvate and its role in the C4 dicarboxylic acid pathway of photosynthesis". The Biochemical Journal. 106 (1): 141–6. doi:10.1042/bj1060141. PMC 1198479. PMID 4305612.
- Reeves RE (June 1968). "A new enzyme with the glycolytic function of pyruvate kinase". The Journal of Biological Chemistry. 243 (11): 3202–4. doi:10.1016/S0021-9258(18)93395-8. PMID 4297474.
- Reeves RE (November 1971). "Pyruvate,phosphate dikinase from Bacteroides symbiosus". The Biochemical Journal. 125 (2): 531–9. doi:10.1042/bj1250531. PMC 1178089. PMID 5144757.
- Reeves RE, Menzies RA, Hsu DS (October 1968). "The pyruvate-phosphate dikinase reaction. The fate of phosphate and the equilibrium". The Journal of Biological Chemistry. 243 (20): 5486–91. doi:10.1016/S0021-9258(18)91972-1. PMID 4302788.