| Firefly luciferase | |||||||
|---|---|---|---|---|---|---|---|
 Structure of Photinus pyralis firefly luciferase. | |||||||
| Identifiers | |||||||
| Organism | |||||||
| Symbol | N/A | ||||||
| PDB | 1LCI | ||||||
| UniProt | P08659 | ||||||
| |||||||
| Firefly luciferase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 1.13.12.7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Firefly luciferase is the light-emitting enzyme responsible for the bioluminescence of fireflies and click beetles. The enzyme catalyses the oxidation of firefly luciferin, requiring oxygen and ATP. Because of the requirement of ATP, firefly luciferases have been used extensively in biotechnology.
Mechanism of reaction
The chemical reaction catalyzed by firefly luciferase takes place in two steps:
- luciferin + ATP → luciferyl adenylate + PPi
- luciferyl adenylate + O2 → oxyluciferin + AMP + light
Light is produced because the reaction forms oxyluciferin in an electronically excited state. The reaction releases a photon of light as oxyluciferin goes back to the ground state.
Luciferyl adenylate can additionally participate in a side reaction with O2 to form hydrogen peroxide and dehydroluciferyl-AMP. About 20% of the luciferyl adenylate intermediate is oxidized in this pathway.[1]
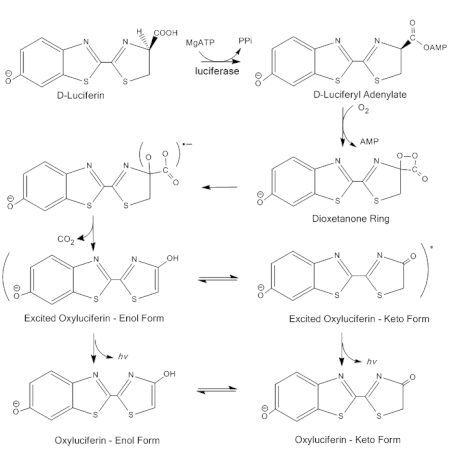
Firefly luciferase generates light from luciferin in a multistep process. First, D-luciferin is adenylated by MgATP to form luciferyl adenylate and pyrophosphate. After activation by ATP, luciferyl adenylate is oxidized by molecular oxygen to form a dioxetanone ring. A decarboxylation reaction forms an excited state of oxyluciferin, which tautomerizes between the keto-enol form. The reaction finally emits light as oxyluciferin returns to the ground state.[2]
Bifunctionality
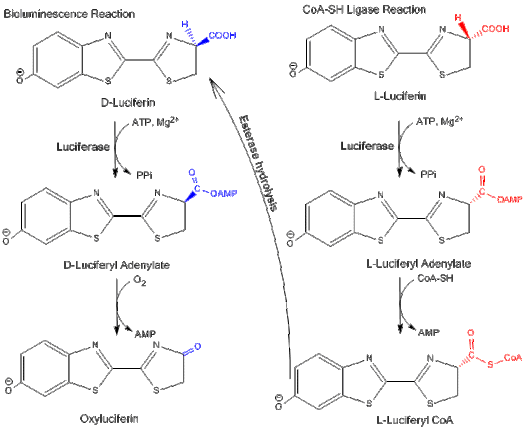
Luciferase can function in two different pathways: a bioluminescence pathway and a CoA-ligase pathway.[4] In both pathways, luciferase initially catalyzes an adenylation reaction with MgATP. However, in the CoA-ligase pathway, CoA can displace AMP to form luciferyl CoA.
Fatty acyl-CoA synthetase similarly activates fatty acids with ATP, followed by displacement of AMP with CoA. Because of their similar activities, luciferase is able to replace fatty acyl-CoA synthetase and convert long-chain fatty acids into fatty-acyl CoA for beta oxidation.[4]
Structure
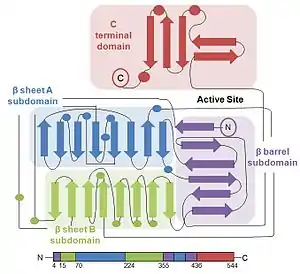
The protein structure of firefly luciferase consists of two compact domains: the N-terminal domain and the C-terminal domain. The N-terminal domain is composed of two β-sheets in an αβαβα structure and a β barrel. The two β-sheets stack on top of each other, with the β-barrel covering the end of the sheets.[2]
The C-terminal domain is connected to the N-terminal domain by a flexible hinge, which can separate the two domains. The amino acid sequences on the surface of the two domains facing each other are conserved in bacterial and firefly luciferase, thereby strongly suggesting that the active site is located in the cleft between the domains.[5]
During a reaction, luciferase has a conformational change and goes into a "closed" form with the two domains coming together to enclose the substrate. This ensures that water is excluded from the reaction and does not hydrolyze ATP or the electronically excited product.[5]
Spectral differences in bioluminescence
Firefly luciferase bioluminescence color can vary between yellow-green (λmax = 550 nm) to red (λmax = 620).[6] There are currently several different mechanisms describing how the structure of luciferase affects the emission spectrum of the photon and effectively the color of light emitted.
One mechanism proposes that the color of the emitted light depends on whether the product is in the keto or enol form. The mechanism suggests that red light is emitted from the keto form of oxyluciferin, while green light is emitted from the enol form of oxyluciferin.[7][8] However, 5,5-dimethyloxyluciferin emits green light even though it is constricted to the keto form because it cannot tautomerize.[9]
Another mechanism proposes that twisting the angle between benzothiazole and thiazole rings in oxyluciferin determines the color of bioluminescence. This explanation proposes that a planar form with an angle of 0° between the two rings corresponds to a higher energy state and emits a higher-energy green light, whereas an angle of 90° puts the structure in a lower energy state and emits a lower-energy red light.[10]
The most recent explanation for the bioluminescence color examines the microenvironment of the excited oxyluciferin. Studies suggest that the interactions between the excited state product and nearby residues can force the oxyluciferin into an even higher energy form, which results in the emission of green light. For example, Arg 218 has electrostatic interactions with other nearby residues, restricting oxyluciferin from tautomerizing to the enol form.[11] Similarly, other results have indicated that the microenvironment of luciferase can force oxyluciferin into a more rigid, high-energy structure, forcing it to emit a high-energy green light.[12]
Regulation
D-luciferin is the substrate for firefly luciferase's bioluminescence reaction, while L-luciferin is the substrate for luciferyl-CoA synthetase activity. Both reactions are inhibited by the substrate's enantiomer: L-luciferin and D-luciferin inhibit the bioluminescence pathway and the CoA-ligase pathway, respectively.[3] This shows that luciferase can differentiate between the isomers of the luciferin structure.
L-luciferin is able to emit a weak light even though it is a competitive inhibitor of D-luciferin and the bioluminescence pathway.[13] Light is emitted because the CoA synthesis pathway can be converted to the bioluminescence reaction by hydrolyzing the final product via an esterase back to D-luciferin.[3]
Luciferase activity is additionally inhibited by oxyluciferin[14] and allosterically activated by ATP. When ATP binds to the enzyme's two allosteric sites, luciferase's affinity to bind ATP in its active site increases.[6]
Homology
Firefly luciferase is thought to be a homolog of long-chain fatty acyl-CoA synthetase because of its ability to synthesize luciferyl-CoA from CoA and dehydroluciferyl-AMP. Inouye tested this hypothesis in 2010 by expressing the cDNA of Photinus pyralis and Lychocoriolaus lateralis luciferses in E. coli through cold shock gene expression.[15] The resulting enzymes were then exposed to long-chain fatty acids, short-chain fatty acids, amino acids, and imino acids. Unsurprisingly, Inouye found that the luciferases only showed adenylation activity when exposed to long-chain fatty acids.
The gene product of CG6178 in Drosophila was also found to have high amino acid sequence similarity with firefly luciferase. While it did show high adenyltation activity when exposed to long-chain fatty acids, there was no luminescence when exposed to oxygen and LH2-AMP– further suggesting that luciferase emerged as a long-chain fatty acyl-CoA homolog due to gene duplication.
Evolution
Phylogenetic analyses performed by Zhang et al. (2020) suggest that the luciferses of the Lampyridae, Rhagopthalmidae, and Phenogodidae families diverged from the Elateridae family 205 Mya.[16] According to phylogenetic data, the emergences of these two luciferases appeared even before the families could diverge– indicating their analogous nature due phenotypic convergences.
See also
References
- ↑ Fraga H, Fernandes D, Novotny J, Fontes R, Esteves da Silva JC (June 2006). "Firefly luciferase produces hydrogen peroxide as a coproduct in dehydroluciferyl adenylate formation". ChemBioChem. 7 (6): 929–935. doi:10.1002/cbic.200500443. PMID 16642538. S2CID 33899357.
- 1 2 3 Baldwin TO (March 1996). "Firefly luciferase: the structure is known, but the mystery remains". Structure. 4 (3): 223–228. doi:10.1016/S0969-2126(96)00026-3. PMID 8805542.
- 1 2 3 Nakamura M, Maki S, Amano Y, Ohkita Y, Niwa K, Hirano T, et al. (June 2005). "Firefly luciferase exhibits bimodal action depending on the luciferin chirality". Biochemical and Biophysical Research Communications. 331 (2): 471–475. doi:10.1016/j.bbrc.2005.03.202. PMID 15850783.
- 1 2 Oba Y, Ojika M, Inouye S (April 2003). "Firefly luciferase is a bifunctional enzyme: ATP-dependent monooxygenase and a long chain fatty acyl-CoA synthetase". FEBS Letters. 540 (1–3): 251–254. doi:10.1016/S0014-5793(03)00272-2. PMID 12681517. S2CID 12075190.
- 1 2 3 Conti E, Franks NP, Brick P (March 1996). "Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes". Structure. 4 (3): 287–298. doi:10.1016/S0969-2126(96)00033-0. PMID 8805533.
- 1 2 Ugarova NN (July 1989). "Luciferase of Luciola mingrelica fireflies. Kinetics and regulation mechanism". Journal of Bioluminescence and Chemiluminescence. 4 (1): 406–418. doi:10.1002/bio.1170040155. PMID 2801227.
- ↑ White EH, Rapaport E, Hopkins TA, Seliger HH (April 1969). "Chemi- and bioluminescence of firefly luciferin". Journal of the American Chemical Society. 91 (8): 2178–2180. doi:10.1021/ja01036a093. PMID 5784183.
- ↑ Fraga H (February 2008). "Firefly luminescence: a historical perspective and recent developments". Photochemical & Photobiological Sciences. 7 (2): 146–158. doi:10.1039/b719181b. PMID 18264582.
- ↑ Branchini BR, Southworth TL, Murtiashaw MH, Magyar RA, Gonzalez SA, Ruggiero MC, Stroh JG (June 2004). "An alternative mechanism of bioluminescence color determination in firefly luciferase". Biochemistry. 43 (23): 7255–7262. doi:10.1021/bi036175d. PMID 15182171.
- ↑ McCapra F, Gilfoyle DJ, Young DW, et al. (1994). Bioluminescence and Chemiluminescence: Fundamentals and Applied.
- ↑ Nakatani N, Hasegawa JY, Nakatsuji H (July 2007). "Red light in chemiluminescence and yellow-green light in bioluminescence: color-tuning mechanism of firefly, Photinus pyralis, studied by the symmetry-adapted cluster-configuration interaction method". Journal of the American Chemical Society. 129 (28): 8756–8765. doi:10.1021/ja0611691. PMID 17585760.
- ↑ Nakamura M, Niwa K, Maki S, et al. (December 2006). "Construction of a new firefly bioluminescence system using L-luciferin as substrate". Anal. Biochem. 47 (1): 1197–1200. doi:10.1016/j.tetlet.2005.12.033.
- ↑ Lembert N (July 1996). "Firefly luciferase can use L-luciferin to produce light". The Biochemical Journal. 317 (1): 273–277. doi:10.1042/bj3170273. PMC 1217473. PMID 8694774.
- ↑ Ribeiro C, Esteves da Silva JC (September 2008). "Kinetics of inhibition of firefly luciferase by oxyluciferin and dehydroluciferyl-adenylate". Photochemical & Photobiological Sciences. 7 (9): 1085–1090. doi:10.1039/b809935a. PMID 18754056. S2CID 2983297.
- ↑ Inouye S (February 2010). "Firefly luciferase: an adenylate-forming enzyme for multicatalytic functions". Cellular and Molecular Life Sciences. 67 (3): 387–404. doi:10.1007/s00018-009-0170-8. PMID 19859663. S2CID 31033141.
- ↑ Zhang R, He J, Dong Z, Liu G, Yin Y, Zhang X, et al. (September 2020). "Genomic and experimental data provide new insights into luciferin biosynthesis and bioluminescence evolution in fireflies". Scientific Reports. 10 (1): 15882. Bibcode:2020NatSR..1015882Z. doi:10.1038/s41598-020-72900-z. PMC 7522259. PMID 32985577.