TaqMan probes are hydrolysis probes that are designed to increase the specificity of quantitative PCR. The method was first reported in 1991 by researcher Kary Mullis at Cetus Corporation,[1] and the technology was subsequently developed by Hoffmann-La Roche for diagnostic assays and by Applied Biosystems (now part of Thermo Fisher Scientific) for research applications.
The TaqMan probe principle relies on the 5´–3´ exonuclease activity of Taq polymerase to cleave a dual-labeled probe during hybridization to the complementary target sequence and fluorophore-based detection.[2] As in other quantitative PCR methods, the resulting fluorescence signal permits quantitative measurements of the accumulation of the product during the exponential stages of the PCR; however, the TaqMan probe significantly increases the specificity of the detection. TaqMan probes were named after the videogame Pac-Man (Taq Polymerase + PacMan = TaqMan) as its mechanism is based on the Pac-Man principle.[3]
Principle
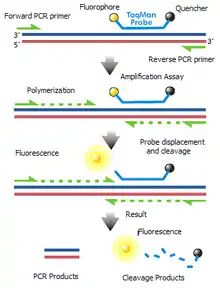
TaqMan probes consist of a fluorophore covalently attached to the 5’-end of the oligonucleotide probe and a quencher at the 3’-end.[4] Several different fluorophores (e.g. 6-carboxyfluorescein, acronym: FAM, or tetrachlorofluorescein, acronym: TET) and quenchers (e.g. tetramethylrhodamine, acronym: TAMRA) are available.[5] The quencher molecule quenches the fluorescence emitted by the fluorophore when excited by the cycler’s light source via Förster resonance energy transfer (FRET).[6] As long as the fluorophore and the quencher are in proximity, quenching inhibits any fluorescence signals.
TaqMan probes are designed such that they anneal within a DNA region amplified by a specific set of primers. (Unlike the diagram, the probe binds to single stranded DNA.) TaqMan probes can be conjugated to a minor groove binder (MGB) moiety, dihydrocyclopyrroloindole tripeptide (DPI3), in order to increase its binding affinity to the target sequence; MGB-conjugated probes have a higher melting temperature (Tm) due to increased stabilization of van der Waals forces. As the Taq polymerase extends the primer and synthesizes the nascent strand (from the single-stranded template), the 5' to 3' exonuclease activity of the Taq polymerase degrades the probe that has annealed to the template. Degradation of the probe releases the fluorophore from it and breaks the proximity to the quencher, thus relieving the quenching effect and allowing fluorescence of the fluorophore. Hence, fluorescence detected in the quantitative PCR thermal cycler is directly proportional to the fluorophore released and the amount of DNA template present in the PCR.
Applications
TaqMan probe-based assays are widely used in quantitative PCR in research and medical laboratories:
- Gene expression assays
- Pharmacogenomics
- Human Leukocyte Antigen (HLA) genotyping
- Determination of viral load in clinical specimens (HIV, Hepatitis)
- Bacterial Identification[7] assays
- DNA quantification
- SNP genotyping
- Verification of microarray results
See also
Notes and references
- ↑ Holland, P. M.; Abramson, R. D.; Watson, R.; Gelfand, D. H. (1991). "Detection of specific polymerase chain reaction product by utilizing the 5'----3' exonuclease activity of Thermus aquaticus DNA polymerase". Proceedings of the National Academy of Sciences of the United States of America. 88 (16): 7276–7280. Bibcode:1991PNAS...88.7276H. doi:10.1073/pnas.88.16.7276. PMC 52277. PMID 1871133.
- ↑ TaqMan Gene Expression - NCBI Projects
- ↑ The Real-Time TaqMan PCR and Applications in Veterinary Medicine - From PacMan to TaqMan - a computer game revisited Archived 2009-05-05 at the Wayback Machine
- ↑ TaqMan Probes: Introduction, functioning and applications
- ↑ Kutyavin IV, Afonina IA, Mills A, Gorn VV, Lukhtanov EA, Belousov ES, Singer MJ, Walburger DK, Lokhov SG, Gall AA, Dempcy R, Reed MW, Meyer RB, Hedgpeth J (2000). "3′-Minor groove binder-DNA probes increase sequence specificity at PCR extension temperatures". Nucleic Acids Res. 28 (2): 655–661. doi:10.1093/nar/28.2.655. PMC 102528. PMID 10606668.
- ↑ Bustin SA (October 2000). "Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays". J. Mol. Endocrinol. 25 (2): 169–93. doi:10.1677/jme.0.0250169. PMID 11013345.
- ↑ AlleleID - Assay Design for Bacterial Identification
External links
1. TaqMan RT-PCR resources- primer databases, software, protocols Archived 2010-11-25 at the Wayback Machine
2. Beacon Designer - Software to design real time PCR primers and probes including SYBR Green primers, TaqMan Probes, Molecular Beacons.