| Phosphatidate phosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 3.1.3.4 | ||||||||
| CAS no. | 9025-77-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
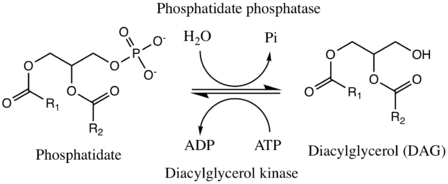
The enzyme phosphatidate phosphatase (PAP, EC 3.1.3.4) is a key regulatory enzyme in lipid metabolism, catalyzing the conversion of phosphatidate to diacylglycerol:[1][2]
- a 1,2-diacylglycerol 3-phosphate + H2O a 1,2-diacyl-sn-glycerol + phosphate
The reverse conversion is catalyzed by the enzyme diacylglycerol kinase, which replaces the hydroxyl group on diacylgylcerol with a phosphate from ATP, generating ADP in the process.
In yeast, the forward direction is Mg2+-dependent, while the reverse process is Ca2+-dependent.[3] PAP1, a cytosolic phosphatidate phosphatase found in the lung, is also -dependent, but PAP2, a six-transmembrane-domain integral protein found in the plasma membrane, is not.[4][5]
Role in the regulation of lipid flux
Phosphatidate phosphatase regulates lipid metabolism in several ways. In short, it is a key player in controlling the overall flux of triacylglycerols to phospholipids and vice versa, also exerting control through the generation and degradation of lipid-signaling molecules related to phosphatidate.[4] When the phosphatase is active, diacylglycerols formed by it can go on to form any of several products, including phosphatidylethanolamine, phosphatidylcholine, phosphatidylserine, and triacylglycerol.[6] Phospholipids can be formed from diacylglycerol through reaction with activated alcohols, and triacylglycerols can be formed from diacylglycerols through reaction with fatty acyl CoA molecules. When phosphatidate phosphatase is inactive, diacylglycerol kinase catalyzes the reverse conversion, allowing phosphatidate to accumulate as it brings down diacylglycerol levels. Phosphatidate can then be converted into an activated form, CDP-diacylglycerol by liberation of a pyrophosphate from a CTP molecule, or into cardiolipin. This is a principal precursor used by the body in phospholipid synthesis. Furthermore, because both phosphatidate and diacylglycerol function as secondary messengers, phosphatidate phosphatase is able to exert extensive and intricate control of lipid metabolism far beyond its local effect on phopshatidate and diacylglycerol concentrations and the resulting effect on the direction of lipid flux as outlined above.[7]
Enzyme regulation
Phosphatidate phosphatase is up-regulated by CDP-diacylglycerol, phosphatidylinositol (formed from reaction of CDP-diacylglycerol with inositol), and cardiolipin. It is down-regulated by sphingosine and dihydrosphingosine. This makes sense in the context of the discussion above. Namely, a build up of products that are formed from phosphatidate serves to up-regulate the phosphatase, the enzyme that consumes phosphatidate, thereby acting as a signal that phosphatidate is in abundance and causing its consumption. At the same time, a build up of products that are formed from DAG serves to down regulate the enzyme that forms diacylglycerol, thereby acting as a signal that this is in abundance and its production should be slowed.[7]
Classification
PAP belongs to the family of enzymes known as hydrolases, and more specifically to the hydrolases that act on phosphoric monoester bonds. This enzyme participates in 4 metabolic pathways: glycerolipid, glycerophospholipid, ether lipid, and sphingolipid metabolism.
Nomenclature
The systematic name is diacylglycerol-3-phosphate phosphohydrolase.[8] Other names in common use include:
- phosphatidic acid phosphatase (PAP),
- 3-sn-phosphatidate phosphohydrolase,
- acid phosphatidyl phosphatase,
- phosphatidic acid phosphohydrolase,
- phosphatidate phosphohydrolase, and
- lipid phosphate phosphohydrolase (LPP).
Types
There are several different genes that code for phosphatidate phosphatases. They fall into one of two types (type I and type II), depending on their cellular localization and substrate specificity.[9]
Type I
Type I phosphatidate phosphatases are soluble enzymes that can associate to membranes. They are found mainly in the cytosol and the nucleus. Encoded for by a group of genes named Lipin, they are substrate specific only to phosphatidate. There are speculated to be involved in the de novo synthesis of glycerolipids.
Each of the 3 Lipin proteins found in mammals—Lipin1, Lipin2, and Lipin3—has unique tissue expression motifs and distinct physiological functions.[10]
Regulation
Regulation of mammalian Lipin PAP enzymes occurs at the transcriptional level. For example, Lipin1 is induced by glucocorticoids during adipocyte differentiation as well as in cells that are experiencing ER proliferation. Lipin2, on the other hand, is repressed during adipocyte differentiation.[3]
Lipin is phosphorylated in response to insulin in skeletal muscle and adipocytes, linking the physiologic action of insulin to fat cell differentiation. Lipin phosphorylation is inhibited by treatment with rapamycin, suggesting that mTOR controls signal transduction feeding into lipin and may partially explain dyslipidemia resulting from rapamycin therapy.[11]
Type II
Type II phosphatidate phosphatases are transmembrane enzymes found mainly in the plasma membrane. They can dephosphorylate other substrates besides phosphatidate, and therefore are also known as lipid phosphate phosphatases. Their main role is in lipid signaling and in phospholipid head-group remodeling.
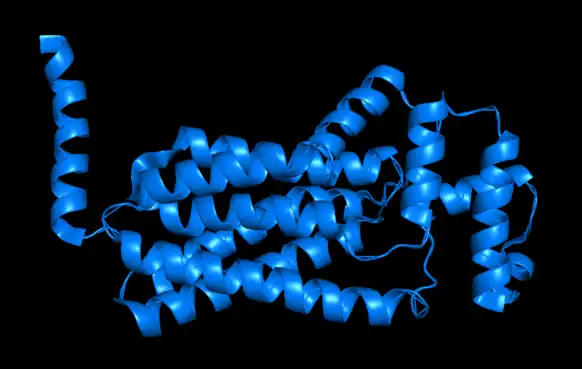
One example of a type II phosphatidate phosphatase is PgpB (PDBe: 5jwy).[12][13] PgpB is one of three integral membrane phosphatases in Escherichia coli that catalyzes the dephosphorylation of phosphatidylglycerol phosphate (PGP) to PG (phosphatidylglycerol).[14] The other two are PgpA and PgpC. While all three catalyze the reaction from PGP to PG, their amino acid sequences are dissimilar and it is predicted that their active sites open to different sides of the cytoplasmic membrane. PG accounts for approximately 20% of the total membrane lipid composition in the inner membrane of bacteria. PgpB is competitively inhibited by phosphatidylethanolamine (PE), a phospholipid formed from DAG. This is therefore an example of negative feedback regulation. The enzyme active site contains a catalytic triad Asp-211, His-207, and His-163 that establishes a charge relay system. However, this catalytic triad is essential for the dephosphorylation of lysophosphatidic acid, phosphatidic acid, and sphingosine-1-phosphate, but is not essential in its entirety for the enzyme's native substrate, phosphatidylglycerol phosphate; His-207 alone is sufficient to hydrolyze PGP.[14]

In the cartoon depiction of PgpB below, one can see its six transmembrane alpha helices, which are here shown horizontally. Of the three PGP phosphatases discussed above, PgpB is the only to have multiple transmembrane alpha helices.[14]
Genes
Human genes that encode phosphatidate phosphatases include:
- PPAP2A (LPP1) – phosphatidic acid phosphatase type 2A
- PPAP2B (LPP3) – phosphatidic acid phosphatase type 2B
- PPAP2C (LPP2) – phosphatidic acid phosphatase type 2C
- PPAPDC1A (PPC1A) – phosphatidic acid phosphatase type 2 domain containing 1A
- PPAPDC1B (PPC1B) – phosphatidic acid phosphatase type 2 domain containing 1B
- PPAPDC2 – phosphatidic acid phosphatase type 2 domain containing 2
- PPAPDC3 – phosphatidic acid phosphatase type 2 domain containing 3
- LPPR2 – lipid phosphate phosphatase-related protein type 2
- LPIN1 – lipin 1[15][16]
- LPIN2 – lipin 2[16]
- LPIN3 – lipin 3[16]
Pathology
Lipin-1 deficiency in mice results in lipodystrophy, insulin resistance, and neuropathy. In humans, variations in Lipin-1 expression levels can result in insulin sensitivity, hypertension, and risk for metabolic syndrome. Serious mutations in Lipin-2 lead to an inflammatory disorder in humans.[10]
References
- ↑ "BRENDA - Information on EC 3.1.3.4 - phosphatidate phosphatase". www.brenda-enzymes.org. Retrieved 2017-03-01.
- ↑ Smith SW, Weiss SB, Kennedy EP (October 1957). "The enzymatic dephosphorylation of phosphatidic acids". J. Biol. Chem. 228 (2): 915–22. PMID 13475370.
- 1 2 Carman, George M.; Han, Gil-Soo (2009-01-30). "Phosphatidic Acid Phosphatase, a Key Enzyme in the Regulation of Lipid Synthesis". The Journal of Biological Chemistry. 284 (5): 2593–2597. doi:10.1074/jbc.R800059200. ISSN 0021-9258. PMC 2631973. PMID 18812320.
- 1 2 Carman, George M.; Han, Gil-Soo (2006-12-01). "Roles of phosphatidate phosphatase enzymes in lipid metabolism". Trends in Biochemical Sciences. 31 (12): 694–699. doi:10.1016/j.tibs.2006.10.003. ISSN 0968-0004. PMC 1769311. PMID 17079146.
- ↑ Nanjundan, Meera; Possmayer, Fred (2003-01-01). "Pulmonary phosphatidic acid phosphatase and lipid phosphate phosphohydrolase". American Journal of Physiology. Lung Cellular and Molecular Physiology. 284 (1): L1–23. doi:10.1152/ajplung.00029.2002. ISSN 1040-0605. PMID 12471011.
- ↑ Martin, Ashley; Gomez-Muñoz, Antonio; Jamal, Zahirali; Brindley, David N. (1991-01-01). "[55] Characterization and assay of phosphatidate phosphatase". In Enzymology, BT - Methods in (ed.). Phospholipases. Phospholipases. Vol. 197. Academic Press. pp. 553–563. doi:10.1016/0076-6879(91)97183-Y. ISBN 9780121820985. PMID 2051944.
- 1 2 Berg, Jeremy (2010-12-24). Biochemistry, 7th Edition. New York: W.H. Freeman and Company. pp. 766–767. ISBN 978-1429229364.
- ↑ "EC 3.1.3.4". www.chem.qmul.ac.uk. Retrieved 2017-03-01.
- ↑ Carman GM, Han GS (December 2006). "Roles of phosphatidate phosphatase enzymes in lipid metabolism". Trends Biochem. Sci. 31 (12): 694–9. doi:10.1016/j.tibs.2006.10.003. PMC 1769311. PMID 17079146.
- 1 2 Reue, Karen; Brindley, David N. (2008-12-01). "Thematic Review Series: Glycerolipids. Multiple roles for lipins/phosphatidate phosphatase enzymes in lipid metabolism". Journal of Lipid Research. 49 (12): 2493–2503. doi:10.1194/jlr.R800019-JLR200. ISSN 0022-2275. PMC 2582367. PMID 18791037.
- ↑ Insulin-stimulated phosphorylation of lipin mediated by the mammalian target of rapamycin Todd A. Huffman, Isabelle Mothe-Satney, John C. Lawrence Proceedings of the National Academy of Sciences Jan 2002, 99 (2) 1047-1052
- ↑ Europe, Protein Data Bank in. "PDB 5jwy structure summary ‹ Protein Data Bank in Europe (PDBe) ‹ EMBL-EBI". www.ebi.ac.uk. Retrieved 2017-03-02.
- ↑ Dillon, Deirdre A.; Wu, Wen-I.; Riedel, Bettina; Wissing, Josef B.; Dowhan, William; Carman, George M. (1996-11-29). "The Escherichia coli pgpB Gene Encodes for a Diacylglycerol Pyrophosphate Phosphatase Activity". Journal of Biological Chemistry. 271 (48): 30548–30553. doi:10.1074/jbc.271.48.30548. ISSN 0021-9258. PMID 8940025.
- 1 2 3 Tong, Shuilong; Lin, Yibin; Lu, Shuo; Wang, Meitian; Bogdanov, Mikhail; Zheng, Lei (2016-08-26). "Structural Insight into Substrate Selection and Catalysis of Lipid Phosphate Phosphatase PgpB in the Cell Membrane". The Journal of Biological Chemistry. 291 (35): 18342–18352. doi:10.1074/jbc.M116.737874. ISSN 1083-351X. PMC 5000081. PMID 27405756.
- ↑ Han GS, Wu WI, Carman GM (April 2006). "The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme". J. Biol. Chem. 281 (14): 9210–8. doi:10.1074/jbc.M600425200. PMC 1424669. PMID 16467296.
- 1 2 3 Donkor J, Sariahmetoglu M, Dewald J, Brindley DN, Reue K (February 2007). "Three mammalian lipins act as phosphatidate phosphatases with distinct tissue expression patterns". J. Biol. Chem. 282 (6): 3450–7. doi:10.1074/jbc.M610745200. PMID 17158099.